INTRODUCTION
Salmonella spp. are rod-shaped, Gram-negative bacteria belonging to the family Enterobacteriaceae and are zoonotic pathogens that typically exhibit motility through flagella (1). Salmonella is recognized worldwide as one of the leading causative agents of foodborne diseases and is a representative waterborne and foodborne pathogen that infects humans through contaminated food or water, causing symptoms such as fever, vomiting, diarrhea, and chills (2). In particular, infections in immunocompromised individuals, infants, and the elderly may progress to severe conditions such as bacteremia or septicemia, making Salmonella a pathogen of significant public health concern (3).
Foodborne salmonellosis is frequently reported in meat and meat products, including poultry, pork, and beef (4, 5), with cross-contamination occurring during slaughtering, processing, distribution, and food preparation identified as the primary routes of transmission (6). In recent years, the mass production and wide-scale distribution of food products have increased the likelihood that a single contamination source may lead to large-scale outbreaks (7, 8, 9). Consequently, the importance of continuous surveillance and epidemiological characterization of meat-derived Salmonella has been increasingly emphasized (10). According to statistical data from the Korean Statistical Information Service (KOSIS), the number of food poisoning outbreaks caused by Salmonella in Korea over the three-year period from 2022 to 2024 showed a steady increase, with 44 cases (14.1%) in 2022, 48 cases (13.4%) in 2023, and 58 cases (21.9%) in 2024 (11).
Salmonella is classified into two species and six subspecies based on antigenic composition and is further divided into approximately 2,600 serotypes according to the specific antigenic properties of the somatic (O), flagellar (H), and capsular (Vi) antigens (12). Serotyping is an important tool for understanding the epidemiological characteristics of Salmonella and tracing sources of infection. Certain serotypes are closely associated with specific food types and host species, and Salmonellaenterica serovar Enteritidis and S. Typhimurium have been reported as the most frequently isolated serotypes in human infections associated with meat, particularly poultry products (13).
Meanwhile, the emergence and spread of antimicrobial-resistant Salmonella have become a global concern due to the use of antibiotics in livestock production (14). When antimicrobial-resistant Salmonella strains are present in food, particularly meat products, transmission to humans may result in treatment failure and prolonged infection duration, thereby substantially increasing public health risks (15). Therefore, understanding the antimicrobial resistance patterns of foodborne Salmonella is critically important for food safety management and antimicrobial resistance surveillance. This study aimed to analyze the antigenic characteristics, including serotypes, and to investigate the antimicrobial resistance profiles of Salmonella isolates recovered from meat products distributed in Gwangju Metropolitan City, thereby providing fundamental data for the prevention and management of foodborne salmonellosis.
MATERIALS AND METHODS
Sample Collection
From January 2022 to December 2024, a total of 994 meat samples were purchased from large discount supermarkets, wholesale food markets, and butcher shops in Gwangju Metropolitan City, Republic of Korea. The samples consisted of carcass and retail cuts of beef, pork, chicken, and duck. After purchase, all samples were immediately transported under refrigerated conditions and tested within 4 h.
Enrichment and Isolation of Salmonella spp.
The enrichment and isolation of Salmonella spp. were performed in accordance with the standard methods for foodborne pathogen investigations and the Korean Food Code (16, 17). Briefly, 25 g of each sample was homogenized with 225 mL of buffered peptone water (Oxoid, Basingstoke, UK) using a Stomacher (ES/Masticator, IUL S.A., Barcelona, Spain) and incubated aerobically at 36°C for 18–24 h. Subsequently, 1 mL and 0.1 mL of the pre-enrichment culture were transferred into tetrathionate broth (Microgiene, Suwon, Korea) and Rappaport–Vassiliadis (RV) broth (Microgiene), respectively, and incubated at 36°C and 42°C for 20–24 h. Each enrichment culture was streaked onto XLD and BG sulfa agar plates (BNF KOREA, Gimpo, Korea) and incubated at 36°C for 20–24 h. Presumptive Salmonella colonies were subcultured onto tryptic soy agar (Oxoid) and incubated at 36°C for 20–24 h, after which final identification was performed using the VITEK® 2 system (bioMérieux, Marcy-l’Étoile, France).
Serotyping
Serotyping of Salmonella isolates was conducted according to the Kauffmann–White scheme using slide and tube agglutination methods (18). Somatic (O) antigens of each isolate were initially identified by slide agglutination using O antisera (Difco, Becton, Dickinson and Company, Sparks, MD, USA). Isolates showing positive reactions were confirmed as Salmonella, and further serogrouping and serotype determination were performed using polyvalent and monovalent O and flagellar (H) antisera (Difco). The specific serotype of each isolate was determined based on the combined O and H antigenic profiles.
Antimicrobial Susceptibility Testing
Antimicrobial susceptibility testing of the isolated Salmonella strains was conducted by broth microdilution method according to the manufacturer’s instructions (bioMérieux). Briefly, isolates were suspended in 3 mL of 0.45% saline in a 5-mL tube and adjusted to a turbidity of 0.6 McFarland using a DENSICHECK (bioMérieux). Subsequently, 145 µL of the suspension was transferred into another 5-mL tube containing 3 mL of 0.45% saline and mixed thoroughly. An AST-N415 card was inoculated with the prepared suspension and analyzed using the VITEK® system. Susceptibility results were interpreted as susceptible, intermediate, or resistant according to the Clinical and Laboratory Standards Institute (CLSI) guidelines (19). The 16 antimicrobial agents tested were ampicillin (AM), amoxicillin/clavulanic acid (AMC), piperacillin/ tazobactam (TZP), cefazolin (CZ), cefotaxime (CTX), ceftazidime (CAZ), cefepime (FEP), ertapenem (ETP), imipenem (IMP), meropenem (MEM), amikacin (AN), gentamicin (GM), tobramycin (TM), ciprofloxacin (CIP), nitrofurantoin (FT), and trimethoprim/sulfamethoxazole (SXT).
RESULT
Isolation of Salmonella spp.
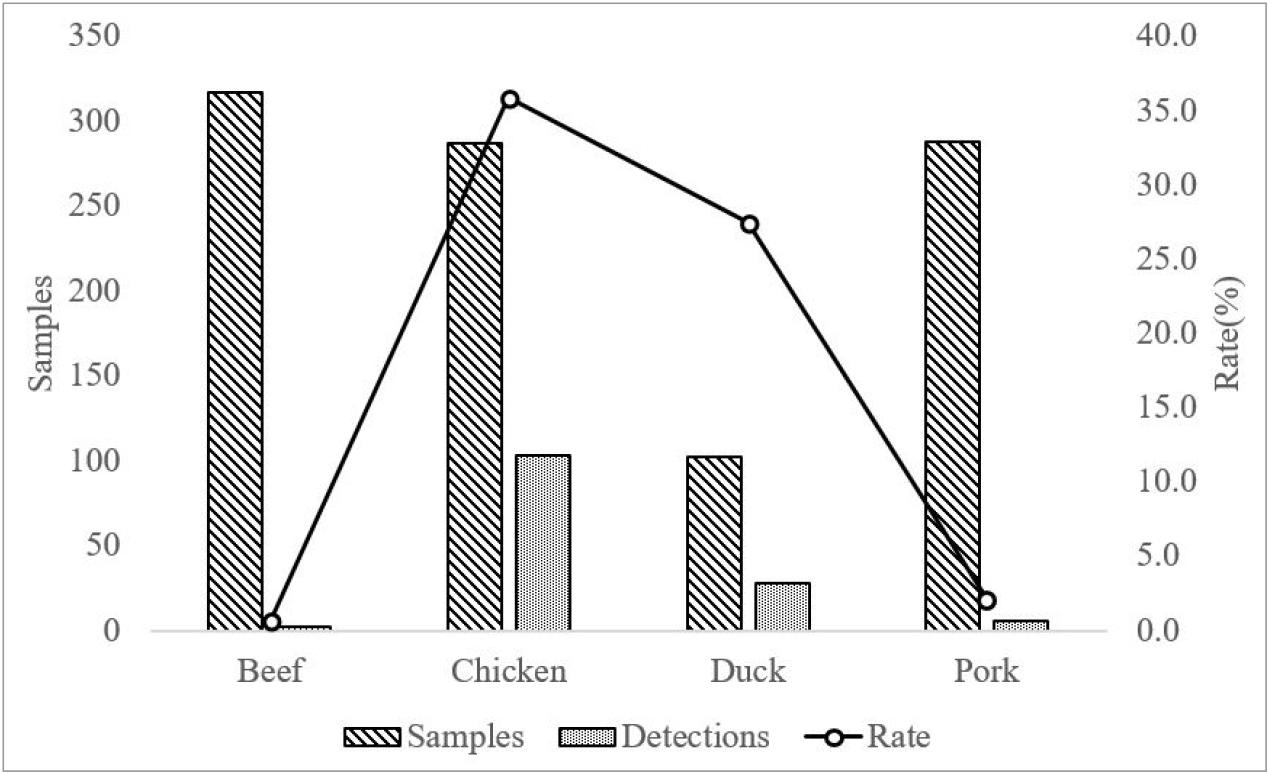
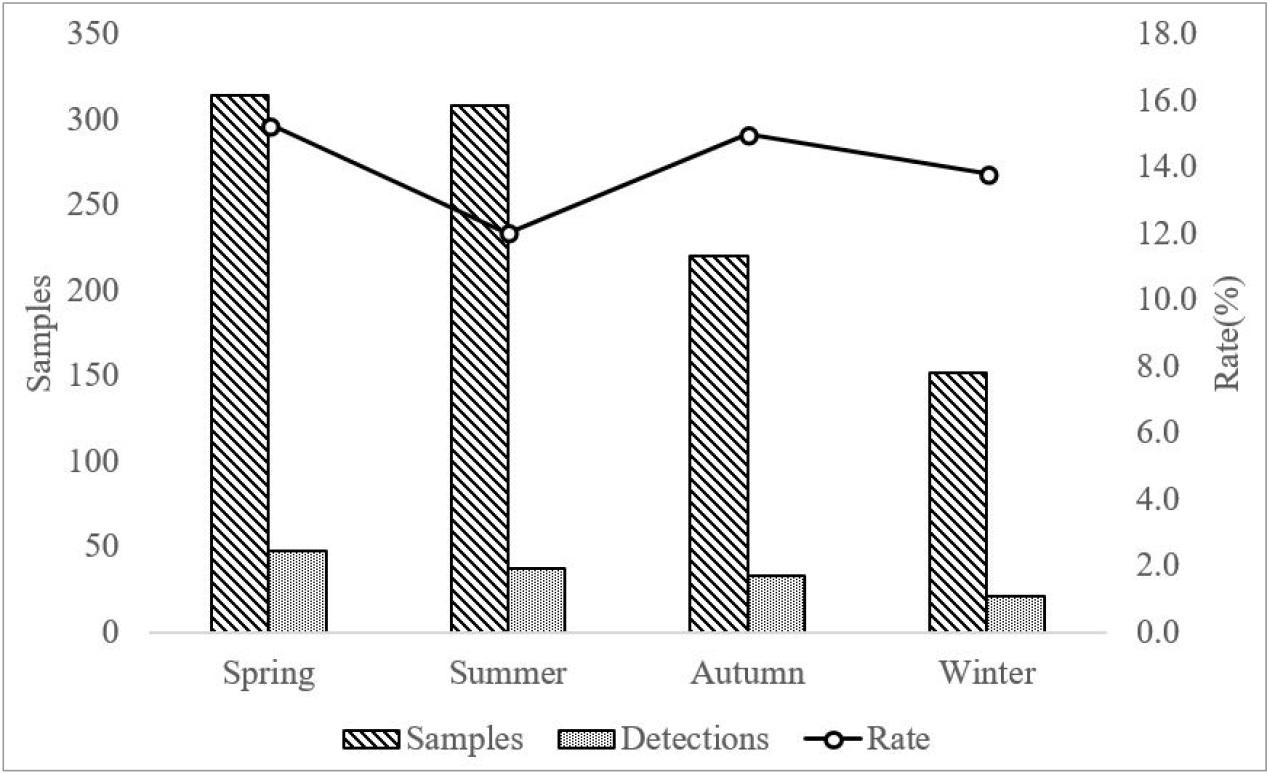
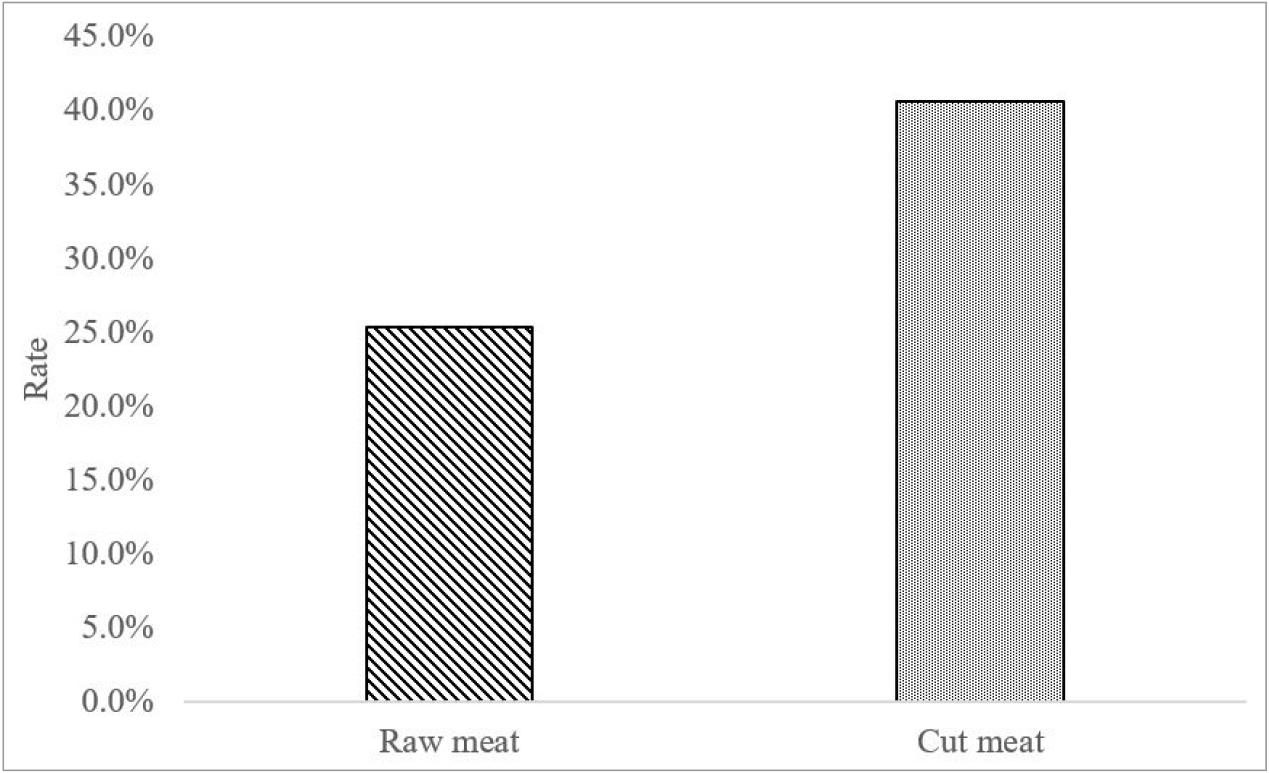
From January 2022 to December 2024, a total of 139 Salmonella isolates were recovered from 994 meat samples distributed in the Gwangju region. By meat type, Salmonella was isolated from 2 of 317 beef samples (0.6%), 103 of 287 chicken samples (35.9%), 28 of 102 duck samples (27.5%), and 6 of 288 pork samples (2.1%) (Fig. 1). Seasonally, Salmonella was isolated from 48 of 314 samples in spring (15.3%), 37 of 308 samples in summer (12.0%), 33 of 220 samples in autumn (15.0%), and 21 of 152 samples in winter (13.8%) (Fig. 2). Among poultry products (chicken and duck), which showed the highest isolation rates, the prevalence of Salmonella differed by processing status; Salmonella was isolated from 25.4% (n = 45) of raw meat samples, whereas a significantly higher isolation rate of 40.6% (n = 86) was observed in cut meat products that underwent additional processing (Fig. 3).
Serotype Distribution of Salmonella Isolates
The serotypes of the isolated Salmonella strains were classified into three serogroups, comprising a total of 15 serotypes, as shown in Table 1. The number of serotypes identified by meat type was nine in chicken, five in duck, two in beef, and two in pork. The predominant serotypes were Salmonella Infantis (98 isolates, 70.5%) and S. Typhimurium (16 isolates, 11.5%), together accounting for 82.0% of all Salmonella isolates. Other serotypes identified included S. I 4,[5],12:i– (4 isolates, 2.9%), S. Bareilly (4 isolates, 2.9%), S. Montevideo (4 isolates, 2.9%), S. Rissen (2 isolates, 1.4%), S. Hadar (2 isolates, 1.4%), S. Enteritidis (2 isolates, 1.4%), S. Agona (1 isolate, 0.7%), S. Bradford (1 isolate, 0.7%), S. Indiana (1 isolate, 0.7%), S. Thompson (1 isolate, 0.7%), S. Braenderup (1 isolate, 0.7%), S. Kentucky (1 isolate, 0.7%), and S. Jamaica (1 isolate, 0.7%).
Table 1.
Serotypes of 139 Salmonella isolates from meat in Gwangju Metropolitan City between 2022 and 2024
Antimicrobial Resistance of Salmonella Isolates
The antimicrobial resistance profiles of Salmonella isolates recovered from meat products were summarized in Table 2. Resistance rates were highest for FT (n = 111, 79.9%), followed by AM (n = 103, 74.1%), CZ (n = 102, 73.4%), CTX (n = 102, 73.4%), TM (n = 85, 61.2%), CIP (n = 48, 34.5%), FEP (n = 36, 25.9%), SXT (n = 16, 11.5%), GM (n = 7, 5.0%), and CAZ (n = 5, 3.6%). No resistance was observed in AMC, TZP, ETP, IPM, MEM and AN.
Abbreviations. Ampicillin, AM ; Amoxicillin/Clavulanic Acid AMC; Piperacillin/Tazobactam, TZP; Cefazolin, CZ; Cefotaxime, CTX ; Ceftazidime, CAZ; Cefepime, FEP ; Ertapenem, ETP; Imipenem, IPM; Meropenem, MEM; Amikacin, AN; Gentamicin, GM; Tobramycin, TM; Ciprofloxacin, CIP; Nitrofurantoin, FT; Trimethoprim/sulfamethoxazole, SXT.
Table 2.
Antimicrobial susceptible of 139 Salmonella isolates from meat in Gwangju Metropolitan City between 2022 and 2024
Abbreviations. Ampicillin, AM; Amoxicillin/Clavulanic Acid, AMC; Piperacillin/Tazobactam, TZP; Cefazolin, CZ; Cefotaxime, CTX; Ceftazidime, CAZ; Cefepime, FEP; Ertapenem, ETP; Imipenem, IPM; Meropenem, MEM; Amikacin, AN; Gentamicin, GM; Tobramycin, TM; Ciprofloxacin, CIP; Nitrofurantoin, FT; Trimethoprim/sulfamethoxazole, SXT.
Difference in Antimicrobial Resistance between Serotypes
Differences in antimicrobial resistance profiles among serotypes are presented in Table 3. Among the 98 S. Infantis isolates, resistance rates were 98.0% for FT, 95.9% for AM, CZ, and CTX, 83.7% for TM, 48.0% for CIP, 30.6% for FEP, 14.3% for SXT, and 4.1% for GM. Among S. Typhimurium isolates (n = 16), resistance was observed to FT (68.8%), CZ (6.3%), and CTX (6.3%). All S. I 4,[5],12:i– isolates (n = 4) exhibited resistance to AM, CZ, CTX, and FEP, with additional resistance to CAZ (75.0%), GM (25.0%), TM (25.0%), and SXT (25.0%). Among S. Montevideo isolates (n = 4), resistance was observed to GM (25.0%) and TM (25.0%). S. Enteritidis isolates (n = 2) showed resistance to AM, CZ, CTX, CAZ, FEP, GM, and FT (50.0% in each). All S. Rissen isolates (n = 2) were resistant to AM (100.0%), with one isolate additionally resistant to SXT (50.0%). Single isolates of S. Bradford and S. Kentucky exhibited resistance to multiple antibiotics. S. Bradford showed resistance to AM, CZ, CTX, and TM, whereas S. Kentucky showed resistance to AM, CZ, CTX, CAZ, FEP, and CIP. In contrast, no antimicrobial resistance was observed in isolates of S. Bareilly (n = 4), S. Jamaica (n = 1), S. Agona (n = 1), S. Thompson (n = 1), S. Braenderup (n = 1), or S. Indiana (n = 1).
Table 3.
Antimicrobial resistance of 130 Salmonella isolates from meat in Gwangju Metropolitan City between 2022 and 2024 by serotypes
Abbreviations. Ampicillin, AM; Amoxicillin/Clavulanic Acid, AMC; Piperacillin/Tazobactam, TZP; Cefazolin, CZ; Cefotaxime, CTX; Ceftazidime, CAZ; Cefepime, FEP; Ertapenem, ETP; Imipenem, IPM; Meropenem, MEM; Amikacin, AN; Gentamicin, GM; Tobramycin, TM; Ciprofloxacin, CIP; Nitrofurantoin, FT; Trimethoprim/sulfamethoxazole, SXT.
Analysis of Multidrug Resistance Patterns
A total of 28 distinct antimicrobial resistance patterns were identified among the 139 Salmonella isolates recovered from meat products. Of these, multidrug-resistant (MDR) Salmonella, defined as resistance to two or more antimicrobial agents, accounted for 106 isolates (76.3%). The MDR patterns ranged from resistance to two to eight antimicrobial agents. The most prevalent pattern was AM–CZ–CTX–TM–FT, observed in 31 isolates (22.3%), followed by AM–CZ–CTX–FEP–TM–CIP–FT in 16 isolates (11.5%) and AM–CZ–CTX–TM–CIP–FT in 15 isolates (10.8%). The distribution of MDR patterns among Salmonella serotypes is presented in Table 4. Among S. Infantis isolates, four (4.0%) exhibited the highest level of multidrug resistance, showing resistance to eight antimicrobial agents, with the patterns AM–CZ–CTX–FEP–TM–CIP–FT–SXT and AM–CZ–CTX–GM–TM–CIP–FT–SXT. In addition, one isolate of S. I 4,[5],12:i– (0.7%) exhibited AM–CZ–CTX–FEP–GM–TM–SXT, and one isolate of S. Enteritidis (0.7%) showed resistance pattern AM–CZ–CTX–CAZ–FEP–GM–FT.
Table 4.
Antimicrobial resistance pattern of 130 Salmonella isolates from meat in Gwangju Metropolitan City between 2022 and 2024 by serotypes
Abbreviations. Ampicillin, AM; Amoxicillin/Clavulanic acid, AMC; Piperacillin/Tazobactam, TZP; Cefazolin, CZ; Cefotaxime, CTX; Ceftazidime, CAZ; Cefepime, FEP; Ertapenem, ETP; Imipenem, IPM; Meropenem, MEM; Amikacin, AN; Gentamicin, GM; Tobramycin, TM; Ciprofloxacin, CIP; Nitrofurantoin, FT; Trimethoprim/sulfamethoxazole, SXT
DISCUSSION
Salmonella remains one of the major causes of food poisoning worldwide and places a significant burden on public health (20). As the spread of antibiotic-resistant Salmonella strains in food production systems increases, clinical management is becoming more complex, and the importance of continuous surveillance is being emphasized (21, 22). This study aimed to provide up-to-date epidemiological data from a food chain perspective by analyzing the serotype distribution and antibiotic resistance patterns of Salmonella strains isolated from retail meat products in Gwangju Metropolitan City.
The results of this study showed that Salmonella is isolated much more frequently from poultry products than from pork or beef. Of the 139 isolated strains, S. Infantis was overwhelmingly dominant (82.0%), while S. Typhimurium (11.5%) and S. Enteritidis (1.4%) were found only in duck and chicken samples, respectively. These results suggest that intensive poultry farming systems can create ecological conditions favorable for the persistence and proliferation of specific serotypes (23). High-density farming, communal production facilities, and vertical integration of the poultry supply chain can facilitate the continuous circulation of dominant strains within farm and processing environments (23, 24).
The high prevalence of S. Infantis, which accounts for more than 70% of isolated strains, raises important considerations regarding the epidemiological independence of the sampled strains. Although the study design involved randomly sampling meat from retail stores in the Gwangju region, detailed information on repeated sampling from the same retail store, manufacturer, production batch, or sampling date was not recorded. Therefore, the possibility that some isolated strains were repeatedly isolated from a common source cannot be ruled out. Clarifying these aspects is crucial for accurately interpreting serotype prevalence and determining whether the observed dominance reflects a widespread distribution or local contamination (25). Future investigations incorporating metadata on source, batch, and production lineage, together with molecular typing or whole-genome sequencing, are warranted to assess the genetic relatedness of isolates and strengthen epidemiological inference.
To interpret the dominance of S. Infantis, it is essential to review the results of this study in relation to long-term domestic epidemiological data. A nationwide analysis of foodborne and waterborne salmonellosis from 1998 to 2007, published in the Journal of the Korean Medical Association, identified S. Enteritidis and S. Typhimurium as major serotypes in clinical infections, while S. Infantis was reported as one of the serotypes that was not notably dominant (26). Importantly, while that study focused primarily on clinical isolates, this study investigated strains isolated from commercially available meat. When comparing serotype distributions, these fundamental differences in sampling methods must be carefully considered.
The markedly high prevalence of S. Infantis in poultry products observed here does not necessarily indicate a novel epidemiological emergence. Rather, it may reflect a redistribution of serotype dominance within the food animal host (24, 27). While clinical surveillance data focuses on serotypes associated with symptomatic infections, food chain surveillance captures ecologically successful serotypes within livestock production systems (28). Previous studies have suggested that S. Infantis is relatively less invasive and induces weaker inflammatory responses compared to S. Typhimurium and S. Enteritidis, which may partially explain the low proportion of S. Infantis in clinical isolates despite its high prevalence in food sources (29). Nevertheless, the global increase in S. Infantis-related outbreaks warrants attention, as changes in host adaptation or the acquisition of resistance can impact public health (30, 31).
Various ecological and anthropogenic factors may act in combination on the spread of S. Infantis in poultry (32). Large-scale intensive livestock farming systems provide stable habitats that promote long-term community formation (23). Additionally, selective pressure resulting from vaccination programs targeting S. Typhimurium and S. Enteritidis can create ecological opportunities for non-vaccinated serotypes, a phenomenon known as serotype replacement (24, 33). Although evidence of a direct causal relationship is still lacking in Korea, patterns of serotype change must be interpreted by considering these epidemiological factors. Overall, the results of this study appear to be an extension of existing epidemiological trends rather than an entirely new phenomenon, emphasizing the importance of integrating foodborne and clinical surveillance data.
It is well known that human salmonellosis increases in warmer seasons (34, 35), but this was not statistically significant in this study. These differences reflect differences in the incidence of infection and the prevalence of contamination in retail stores. Modern refrigerated and standardized distribution systems can mitigate contamination levels in retail stores from external temperature fluctuations (36). Furthermore, the observation of much higher isolation rates in cut meat compared to raw meat supports the hypothesis that secondary processing steps such as cutting and mechanical bone removal contribute to cross-contamination (37). These results highlight the need for a stepwise control strategy within the meat processing environment.
From the perspective of antimicrobial resistance, the findings of this study indicate considerable resistance to several antibiotic classes. The World Health Organization (WHO) AWaRe classification classifies antibiotics into Access (first-line treatments with a low risk of resistance), Watch (antibiotics with a high risk of resistance and limited use), and Reserve (antibiotics used as a last resort for multidrug-resistant infections), providing a framework for antimicrobial management (38) According to the National Antimicrobial Monitoring Report, in 2024, penicillin-based antibiotics were the most abundant in South Korea (approximately 284 tons), while sales of key antibiotics, including third and fourth-generation cephalosporins and quinolones, were much lower (approximately 46 tons) (39). According to the WHO AWaRe classification system, high resistance was observed to several Access antibiotics, including ampicillin, cefazolin, cefotaxime, and tobramycin. These aspects suggest that widespread and long-term use of first-line treatments may have exerted cumulative selective pressure within livestock production systems (40, 41). Watch antibiotics such as ceftazidim, cefepime, and ciprofloxacin have been found to be resistant despite relatively low usage, raising concerns that the therapeutic effects of important antibiotics may gradually decline (42). The high resistance rate to nitrofurantoin observed in this study is consistent with previous studies of meat-derived Salmonella enterica, and loss of nitrofuran reductase activity due to meaningless mutations in the snrA and cnr genes has been suggested as a resistance mechanism (43). These results highlight the need to consider both AWaRe classification and regional antibiotic susceptibility aspects in the clinical management of salmonellosis. In regions with high antibiotic resistance, relying solely on Access antibiotics may not guarantee effective treatment.
The Salmonella isolates exhibited a variety of antibiotic resistance patterns depending on the serotype. Multidrug resistance (MDR) was particularly pronounced in S. infantis, with common resistance patterns including β-lactam antibiotics, aminoglycoside antibiotics, quinolone antibiotics, and trimethoprim/sulfamethoxazole combinations. Of these, the AM-CZ-CTX-TM-FT and AM-CZ-CTX-FEP-TM-CIP-FT patterns occurred with considerable frequency. The involvement of third- and fourth-generation cephalosporins and quinolone antibiotics significantly limits treatment options and may hinder empirical treatment strategies (42, 44). The concentration of MDR phenotypes within a single dominant serotype raises concerns about clonal spread and horizontal transmission of poultry production chain resistance determinants (45). These trends are consistent with previous national and international studies of S. infantis, which identified antibiotic exposure and the accumulation of resistance genes during livestock production and distribution as major contributing factors (24, 46, 47, 48).
In particular, the AM-CZ-CTX-TM-FT multidrug resistance pattern was repeatedly observed in S. Infantis isolates. Several genetic mechanisms underlie this multidrug resistance, including plasmids containing a broad range of β-lactamase (ESBL) genes (e.g., the bla_CTX-M family), aminoglycoside resistance gene cassettes, type I integrons, and internationally reported MDR megaplasmids associated with S. Infantis (44, 49). However, this study was limited to phenotypic antimicrobial susceptibility testing and did not include plasmid characterization, ESBL gene detection, integron analysis, or molecular typing. Therefore, it is not possible to determine whether the observed MDR pattern is due to clonal propagation of a single strain or horizontal gene transfer between various strains.
Future research should identify the genetic basis and dynamics of multidrug-resistant S. infantis, including plasmid replication assays, ESBL gene screening, integron characterization, and sequence-based approaches such as MLST or whole-genome sequencing. These analyses are crucial for distinguishing between clonal spread and horizontal acquisition of resistance genes, and for providing information on targeted interventions within poultry production and public health strategies.
The heterogeneity of the observed serotype resistance profiles further suggests that antibiotic resistance in Salmonella is shaped by a complex interaction between genetic background, environmental adaptation, and selective pressures (50). Even within the same species, resistance phenotypes can differ considerably at the serotype and strain levels (49). Therefore, antibiotic management and surveillance strategies should take serotype-specific dynamics into account rather than treating Salmonella as a homogeneous organism.
Overall, this study highlights the evolution of Salmonella ecology during the food production process. The major emergence of multidrug-resistant (MDR) S. Infantis in retail poultry meat highlights the need for integrated “One Health” surveillance linking the animal, food, and human sectors (51). Future studies, including future full-length dielectric sequencing and resistance gene profiling, are essential to determine whether S. Infantis is experiencing continued epidemiological spread in Korea or remains a food-related serotype with relatively limited clinical impact. Enhancing coordinated monitoring at farm and processing stages and implementing customized interventions are critical to mitigating public health risks associated with antibiotic-resistant Salmonella(24, 33, 37).
CONCLUSION
In this study, Salmonella strains were isolated from meat products distributed in the Gwangju region, and the serotypes and antimicrobial resistance profiles of the isolates were analyzed to investigate resistance patterns. The main findings are summarized as follows.
1.Among 994 meat samples collected in the Gwangju region, Salmonella was isolated from 14.0% (n = 139) of the samples. By meat type, the isolation rates were 35.9% (n = 103) in chicken, 27.5% (n = 28) in duck, 2.1% (n = 6) in pork, and 0.6% (n = 2) in beef.
2.Serotyping based on antigen agglutination reactions identified 15 serotypes from the 139 isolated strains. Of these, 5 were type B, 8 were type C, and 2 were type D. S. Infantis (70.5%, n = 98) and S. Typhimurium (11.5%, n = 16) were the major serotypes.
3.Antimicrobial resistance analysis revealed that resistance to FT (79.9%) was the most frequently observed, followed by AM (74.1%), CZ (73.4%), and CTX (73.4%). High resistance rates were particularly observed to penicillin-class and cephalosporin-class antibiotics.
4.A total of 28 antimicrobial resistance patterns were identified among the 139 Salmonella isolates. Multidrug-resistant (MDR) Salmonella, defined as resistance to two or more antimicrobial agents, accounted for 76.3% (n = 106) of the isolates. The MDR patterns ranged from resistance to two to eight antimicrobial agents, with the five-drug resistance pattern AM–CZ–CTX–TM–FT being the most prevalent, observed in 31 isolates (22.3%). Notably, 98.0% (n = 96) of S. Infantis isolates exhibited multidrug resistance.