INTRODUCTION
Klebsiella pneumoniae is a member of the Enterobacteriaceae and is found in various habitats, including water, soil, and the intestines and oropharynx of humans and animals (1, 2). There are two main pathotypes of K. pneumoniae, distinct phenotypically, clinically, and genetically: classical K. pneumoniae (C-K. p) and hypervirulent K. pneumoniae (hv-K. p) (3, 4). Phenotypically, hv-K. p shows hypermucoviscous colonies on an agar plate, which can be demonstrated by the string test (>5mm) (5). Clinically, unlike C-K. p, which generally causes hospital-acquired infections in immunocompromised individuals, hv-K. p is implicated in serious community-acquired infections in young and healthy individuals. Initially, hv-K. p was reported in Taiwan in the 1980s in cases of community-acquired liver abscesses; subsequently, infections have been reported in the Middle East (6, 7, 8), India (9), China (10), Europe (11), and America (12). hv-K.p hepatic infections tend to metastasize to other body sites, a clinical feature not previously known in C-K. p or other Enterobacteriaceae(13) Moreover, many reports worldwide have documented extrahepatic infections caused by hv-K. p, including urinary tract infections, pneumonia, kidney abscesses, and septic arthritis (14, 15). At the genomic level, hv-K.p harboring several virulence-associated genes, including crampA, rampA2, iroB, and iucA within a large virulence plasmid termed pLVPK, enables it to be more virulent than the classical pathotypes. This cardinal difference has been employed to differentiate hv-K.p from C-K.p in clinical samples (16, 17). Most previous studies investigated the prevalence of hv-K. p in human clinical samples (4, 18, 19, 20). However, little is known about the prevalence of hv-K. p in non-human sources. It has been hypothesized that non-clinical sources, particularly animals, represent an important reservoir of diverse clones of K. pneumoniae that contribute to inter-species transmission between humans and animals (21). Poultry and the poultry industry, in particular, provide a thriving ground for propagating and spreading pathogens in communities (22). Studies investigating the occurrence, prevalence, and antibiogram profile of hv-Kp in nonclinical sources might provide valuable information that aids in drawing a complete epidemiological figure of that virulent pathotype of K. pneumoniae. Therefore, this study investigated the occurrence and antibiogram profile of hv-K. p in broiler chickens at chicken slaughter shops, representing the final stage in the poultry industry.
MATERIALS AND METHODS
Study Design, Location, and Sample Collection
This cross-sectional study involved 60 broiler chicken slaughter shops in Thi-Qar province, the fourth-largest province in south Iraq. Broiler chicken slaughter shops, which specialise in slaughtering, de-feathering, and trimming chickens, are widely distributed in Thi-Qar province. We collected 440 broiler chicken cloacal swabs from January to September 2024. The samples were transported via transport medium (Alkaline Peptone Water, HiMedia® laboratories, India) in ice-cooled containers to the microbiology laboratory of Al-Nasiriyah Technical Institute for bacterial isolation and molecular identification.
K. pneumonia Isolation, Identification, and Antibiogram Profiling
The cloacal swabs were directly streaked onto MacConkey agar (HiMedia Laboratories®, India) and incubated at 37°C for 24 h. Colonies that showed a mucoid pink colour on the MacConkey agar plate were further purified by sub-culturing on the same culture media. K. pneumoniae was identified using the automated system VITEK II compact (GN ID card, lot No. 2412933103, bioMérieux, France). Antimicrobial susceptibility testing of K. pneumoniae was conducted using VITEK II system (GN ID card, AST N419; lot No. 0442845204; bioMérieux, France). The following antibiotics were tested: ceftriaxone, cefepime, ciprofloxacin, gentamicin, aztreonam, imipenem, meropenem, tetracycline, and azithromycin. Multi-drug resistant isolates were defined as those that showed resistance against three or more antibiotics of different classes, while isolates that showed resistance against imipenem and/or meropenem were classified as carbapenem- resistant isolates (4).
Phenotypic Detection of Hypermucoviscosity (String Test)
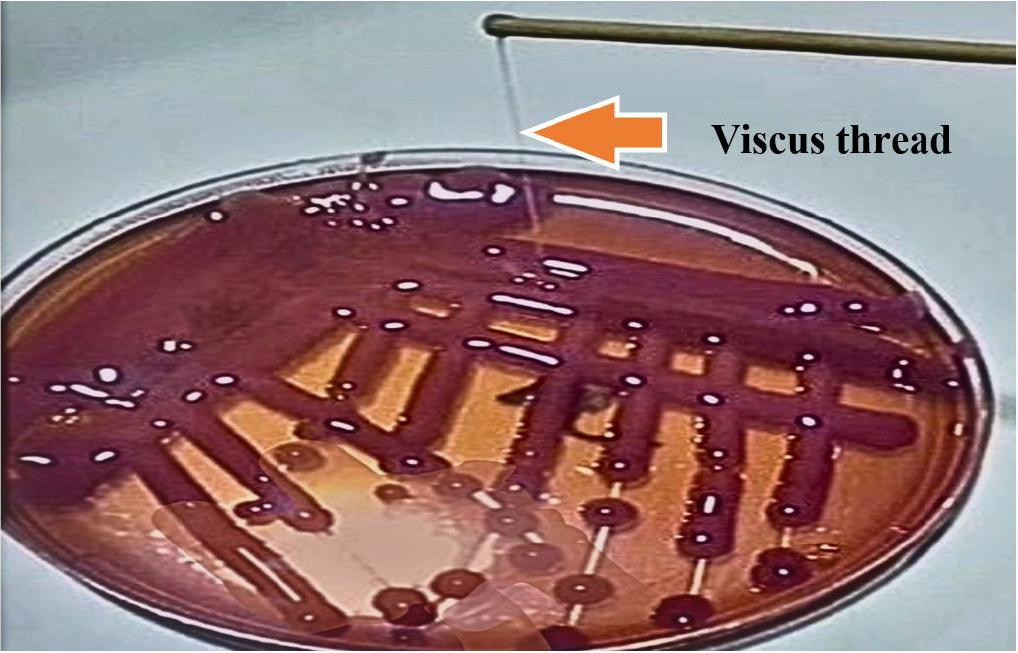
To detect the hypermucoviscosity character of K. pneumoniae, the overnight pure colonies of K. pneumoniae on the blood agar plate were stretched using an inoculation wire loop. The formation of a long, viscous thread (>5 mm) is considered a positive string (18).
Molecular Confirmation of K. Pneumoniae and Detection of hv-K.p Pathotype
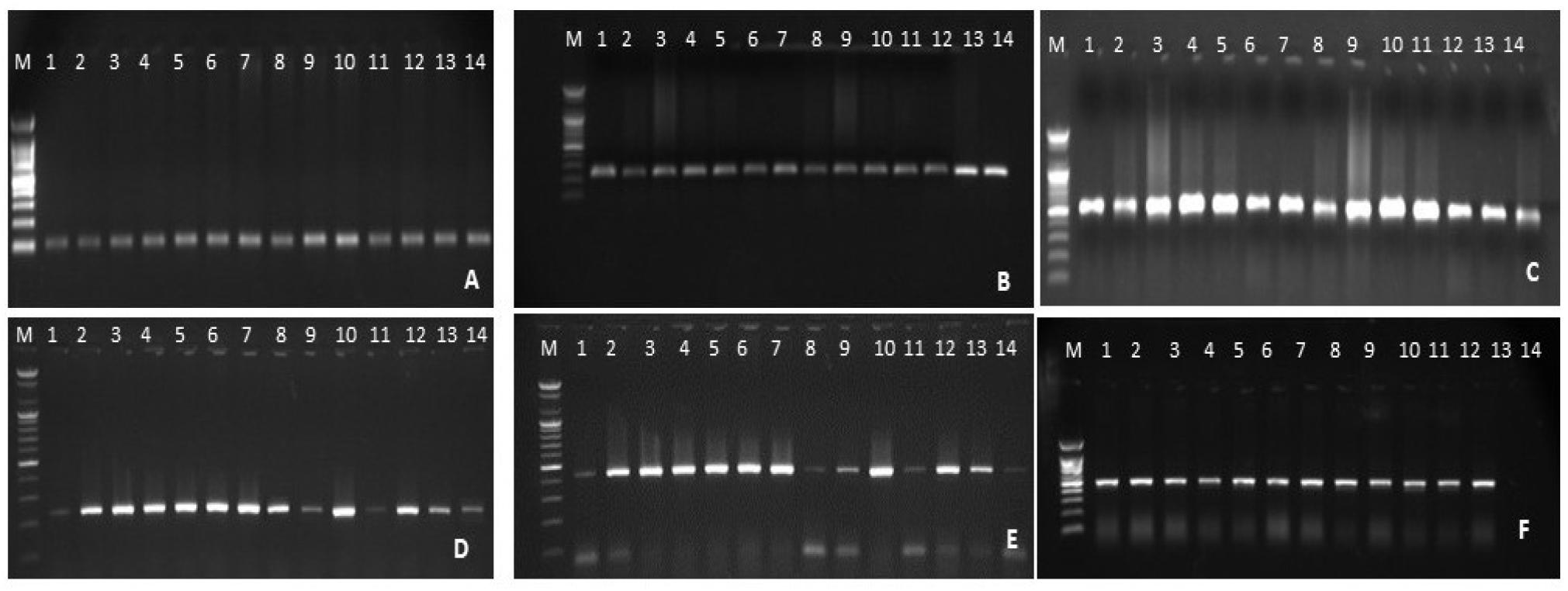
Polymerase chain reaction (PCR) was performed by using six primers Table 1. The first primer targeted the ribosomal DNA internal transcribed spacer (16-23S rDNA ITS) to confirm the identification of K. pneumoniae. The other five primers targeted the hypervirulent-associated genes, namely, iroB, iucA, crmpA, rmpA2, and peg344, as these genes are considered molecular markers that differentiate hypervirulent pathotypes with diagnostic accuracy exceeding 95% (16, 17). The PCR mixture was carried out in 0.2 ml tubes containing 5µl bacterial extracted DNA, 1µl (10 pmol) of each forward and reverse primer, and 5µl of Premix Accuapure (Bioneer, Korea). The remaining volume was completed with deionised water. The thermocycling conditions were as follows: initial denaturation at 94°C, 4 min for the 16-23S rDNA ITS; 95°C, 2 min for the hypervirulent genes, denaturation at 94°C, 45 S for16-23S rDNA ITS; 95°C, 30 S for the five hypervirulent genes, annealing at specified temperatures in the Table 1 for 30 S. Extension at 72°C for 5 min (16-23S rDNA ITS), 30 S (iroB), 50 seconds (iucA), 60 seconds (crmpA), and 40 S (rmpA2 and Peg-344). The final extension temperature was set at 72°C for 10 min. The number of reaction cycles was 30 for 16-23S rDNA ITS, and 25 for the five virulence genes. The amplification products were visualised through electrophoresis using agarose gel (2%) stained with Ethidium bromide. Photographs were taken using the gel documentation system (Atta, Japan).
Table 1.
The primer sequences, product sizes, and their annealing temperatures used in this study
| Gene |
Primer sequence 5'-3' | Amplicon size (bp) |
Annealing temperature | References |
|---|---|---|---|---|
| 16-23S rDNA ITS |
F- ATTTGAAGAGGTTGCAAACGAT R- TTCACTCTGAAGTTTTCTTGTGTTC | 130 | 57°C | (23) |
| iroB |
F- ATCTCATCATCTACCCTCCGCTC R- GGTTCGCCGTCGTTTTCAA | 235 | 59°C | (17) |
| iucA |
F-AATCAATGGCTATTCCCGCTG R-CGCTTCACTTCTTTCACTGACAGG | 239 | 59°C | (17) |
| crmpA |
FGTAATAGAGATATAAATATCATATTGA R- CATCTTTCATCAACCATTTC | 588 | 50°C | (17) |
| rmpA2 |
F- GTGCAATAAGGATGTTACATTA R- GGATGCCCTCCTCCTG | 430 | 50°C | (17) |
| Peg-344 |
F- CTTGAAACTATCCCTCCAGTC R- CCAGCGAAAGAATAACCCC | 508 | 53°C | (17) |
RESULTS
The Recovery Rate of K. pneumoniae from Broiler Chicken Intestine
Of 440 cloacal swabs, only 130 (29.5%) were identified as K. pneumoniae based on phenotypic characteristics and molecular detection of the 16-23S rDNA ITS gene, (Fig. 1).
Molecular Detection of Hypervirulent K. pneumoniae
The molecular detection of hypervirulence-associated genes revealed that 35 (26.9%) of 130 K. peumoniae isolates were positive for the hypervirulent genes with varying frequencies and thus identified as hv-K. p, (Fig. 1). The remaining 95 (73.1%) isolates were negative for all hypervirulence-associated genes and thus were designated as C-K. p. Among the hv-K. p, the most frequently detected gene was iucA, found in 31 of the 35 isolates (88.6%), followed by crmpA in 26 (74.3%), iroB in 22 (62.9%), rmpA2 in 16 (45.7%), and peg-344 in 12 (34.3%).
Comparison Between String Test and Molecular Detection of hv-K.p
Of the 130 K. pneumoniae isolates, only 25 (19.2%) were string positive, (Table 2, Fig. 2). Of the 25 string-positive isolates, 20 (80%) were identified to hv-K. p based on the molecular detection of hypervirulent-associated genes. On the other hand, only 15 (14.3%) of hv-K. p did not show hypermucoviscosity (string negative). Statistical analysis revealed a significant difference (P ≤ 0.05) in diagnosing hv-K. p between the two methods, string test and molecular method. Moreover, compared with the molecular detection of hypervirulence genes as the reference method, the sensitivity, specificity, and positive and negative predictivity values of the string test were 57.1%, 94.7%, 80%, and 85.7%, respectively.
Virulence gene profiling and the Hypermucoviscosity Trait of hv-K.p isolates
The hv-K. p isolates in this study were categorised into five groups (I-V) based on the number of detected hypervirulence-associated genes. Group III, characterized by presence of three hypervirulent genes was the most prevalent, accounting for 14 of the 35 isolates (40%). This was followed by group I and II, which harbour five and four hypervirulent genes respectively; each group comprised 17.1% of the Hv-K. p isolates. Group V, with only one gene, included (14.3%), while group IV, positive for two genes, was the least common, comprising (11.4%) of the isolates. Interestingly, there was a clear correlation between the number of hypervirulence-associated genes and the hypermucoviscosity phenotype as determined by string tests. All isolates in group I exhibited the hypermucoviscosity trait, followed by 80.4% of isolates in group II. In contrast, only 25% of group IV isolates showed hypermucoviscosity, and none of group V isolates exhibited this characteristic. The relationship between virulence genes profile (in terms of number and gene combinations) and the hypermucoviscosity phenotype is summarized in Table 3.
Table 3.
Genotyping of hypervirulent K. pneumoniae isolates in this study based on the number and combination of detected genes and their hypermucoviscosity traits
Antibiotic Resistance Profile of hv-K. p isolates
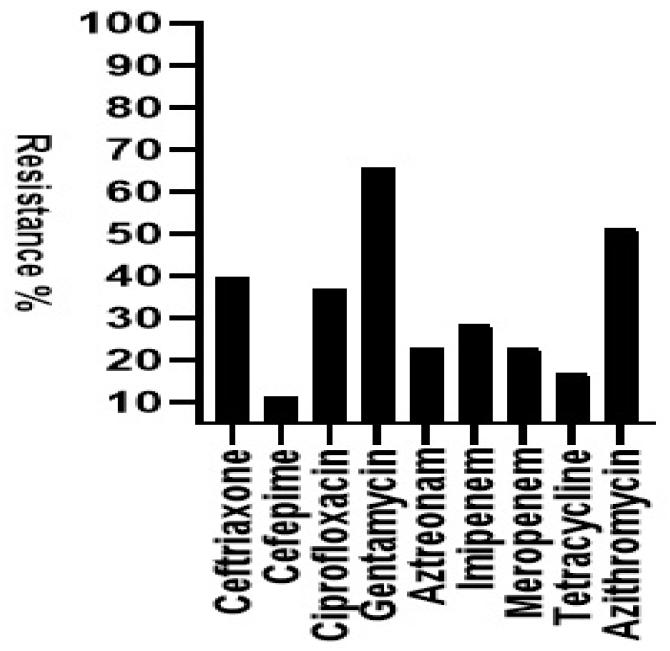
The antibiotic resistance profiles of the 35 hv-K. p isolates are presented in Fig. 3. Overall, the isolates exhibited relatively low resistance rates, not exceeding 70%. The highest resistance was observed against gentamicin, with 23 isolates (65.7%) showing resistance, followed by azithromycin, with 18 (51.4%) isolates. The lowest resistance rate was observed for cefepime, with only four (11.4%) demonstrating resistance. Multi-drug resistance was found in 21 of the 35 isolates (60%). Additionally, carbapenem resistance was detected in 16 isolates (45.7%), including two isolates that were resistant to both imipenem and meropenem.
DISCUSSION
Although four decades have passed since hv-K. p was first recognised in Taiwan, several aspects of its epidemiology remain unexplored (15). One such gap is the potential role of animals as reservoirs contributing to hv-K. p transmission in communities. To our knowledge, this is the first study in Iraq and the Middle East to explore the occurrence of hv-K. pin broiler chickens and highlight chicken slaughter shops as a potential public health risk.
This study revealed a K. pneumoniae prevalence of 29.5% in broiler chickens, consistent with a previous study from northern Iraq (24). This prevalence aligns with similar findings from healthy broiler chickens in Ethiopia, 22% (25), Indonesia, 22.5% (26), and Norway, 25.8% (27). Also, our finding is concordant with the results of a study conducted in Bangladesh, where 34.7% of chicken meat samples were positive (28). However, our prevalence was lower than those reported by Kahin et al. in Ethiopia (29), and Chika et al. in Nigeria (60%) (30). Conversely, our finding exceeded those from Egypt, 15% (31); China, 14.5% (32) and another study in Indonesia, 7.8% (33). This discrepancy may be due to differences in the geographical location, sample types, culturing techniques, and poultry farming practices (25).
The identification of hv-K.p poses a challenge due to limitations of phenotypic methods. However, molecular detection of hypervirulence-associated genes such as crmpA, rmpA2, iroB, iucA, and peg-344, has proven be a highly specific approach with up to 96% accuracy (17). Using this method, we found a 26.9% prevalence of hv-K.p in broiler chickens. Unfortunately, data on hv-K.p. prevalence in chickens are scarce, limiting direct comparison. However, our findings are comparable to a previous study from China (31). Other studies have detected hv-K.p in various animal sources, including oysters, 62.5% (34), pork,16.7% (35), and farm animals, 24.2% (36). Additionally, hypervirulent multidrug-resistant K. pneumoniae has been recorded in captive marmosets in Brazil (37), sea lions in California were identified as a potential zoonotic source (38). Our prevalence also parallels findings in human clinical isolates: Iraq, 43.3% (7); Iran, 32.2% (20), and Sudan, 31.6% (39). However, it was notably higher than that reported in European clinical isolates, including Italy, 3.7% (40) and Germany, 7.6% (41).
Among the hypervirulence-associated genes, the most frequently detected gene in our study were iucA, followed by crmpA, iroB, rmpA2, and peg-344. Our findings regarding the icuA gene, which encodes aerobactin, align with previous studies showing it as a dominant siderophore gene in 90% of hv-K.p strains and rarely present in C-Kp (3, 42). Asian studies have similarly reported iucA prevalence ranging from 60-90% (43). The crmpA gene involved in hypermucoviscosity and virulence (3) was detected in 74.3% of our isolates, consistent with other reports (34, 44). Meanwhile, the peg-344 gene, a metabolic transporter protein, was the least frequently detected, consistent with the report from Sanikhani (20). The variation in gene detection frequencies may be due to incomplete virulence plasmid (pVLK) in some isolates (16).
A statistical significance difference (P≤0.05) was observed between the string test (19.2%) and the molecular method (26.9%) for detecting of hv-K.p, with the string test demonstrate low sensitivity 57.1%. This is consistent with the previous studies that reported string test sensitivity ranging from 50% to 90% (17, 45, 46). In our data 14.3% of the isolates that were negative string tests were positive for hypervirulence genes and five isolates of C-Kp tested positive by string test. These finding reinforce the notion that some hv-K.p strains may falsely appear hypermucoviscosity (18, 19, 47). Moreover, as highlighted by Catalán-Nájera et al. (48) that hypermucoviscosity and hypervirulence are distinct and should not be used interchangeably. The expression of hypermucoviscosity can also be influenced by the growth conditions and the type of agar medium used (42, 49).
Regarding the antibiotic resistance, our finding of relatively low resistance rate align with previous studies, indicating that hv-K.p strains generally exhibit low resistance rate compared to those of the classical pathotypes (43, 44). Non-clinical settings: animals often display lower resistance rate than human clinical samples (50). However, remarkable resistance rates were found against gentamicin, azithromycin, and ceftriaxone, with similar findings also being obtained previous reports (31, 51, 52). The widespread use of gentamicin and erythromycin in poultry farms at high doses likely contributes to this trend (53). In our study, 60% of isolates exhibited multi-drug resistance, which is lower than 100% reported by others (33, 53), and the 88.9% observed in animal isolates (50). This low rate may be due to our isolates being predominantly hypervirulent pathotypes as multi-drug resistant phenotype is generally more prevalent among classical strains pathotypes (4). Notably, we detected a high prevalence (45.7%) carbapenem-resistant hv-K.p (CR-hv-K.p). While, this contrast with some studies (54), it along with other that reported CR-hv-K.p ranging from 31.8%- 55.9% (7, 31, 34), and even a higher rate in China (64.1%) (54). The presence of CR-hv-K.p in chickens strongly suggest human to animals transmission, as carbapenem such as imipenem or meropenem are not used in poultry. Although K. pneumoniae is not traditionally considered a zoonotic threat, its ability to acquire and transfer the resistance and virulence plasmids through horizontal and vertical gene transfer, necessitate reconsideration of it zoonotic potential. A key strength of this study is that it is the first to investigate hv-K.p in chicken within this region, a relevant focus given the high consumption of chicken meat due to its affordability compared to other meats. Moreover, unlike previous studies that depended on string test or screened only one or two hypervirulence- associated genes, we evaluated all five proposed hypervirulence associated genes and compare molecular method to the string test in defining hypermucoviscosity. The relatively large sample size further enhances the reliability of our findings. However, this study has limitations, we were unable to obtain clinical isolate from hospitals preventing the assessment the genetic relatedness between chicken and clinical isolates. moreover, since samples were taken from intestinal swabs of slaughtered chickens, data on the clinical status and demographic characteristics, including age, sex, and breed, were not considered in this study.