INTRODUCTION
Following the COVID-19 pandemic, the epidemiology of respiratory infectious diseases has changed globally. In Korea, the re-emergence of various respiratory infections has been increasingly observed among children and adolescents after the relaxation of public health control measures (1, 2). However, the current national surveillance system for respiratory infectious diseases primarily focuses on viral pathogens. Moreover, region-specific data on bacterial respiratory infections remain relatively limited.
Pertussis is a vaccine-preventable disease whose incidence declined following the introduction of routine immunization; nevertheless, recurrent resurgences have been reported domestically and globally in recent years, particularly among school-aged children and adolescents (3, 4, 5). Waning immunity after vaccination, together with atypical clinical presentations, allow adolescents and adults to act as major reservoirs for transmission to infants, posing a significant public health concern (6, 7). In addition, the potential involvement of Bordetella species closely related to B. pertussis has highlighted limitations in conventional diagnostic and surveillance systems that focus on a single pathogen (8, 9).
Meanwhile, Mycoplasma pneumoniae is a major causative agent of community-acquired pneumonia in children and adolescents, with periodic nationwide epidemics reported in Korea (10, 11). In recent years, the prevalence of macrolide- resistant M. pneumoniae strains has continued to increase, raising concerns regarding treatment failure and severe clinical outcomes (12, 13, 14). In addition, Streptococcus pneumoniae and Haemophilus influenzae remain key bacterial respiratory pathogens in pediatric populations, with persistent challenges related to antimicrobial resistance and to distinguishing colonization from true infection (15, 16).
Multiplex polymerase chain reaction (PCR) assays have been widely adopted in clinical and surveillance settings because they enable the simultaneous detection of multiple respiratory pathogens from a single specimen. Nevertheless, careful interpretation of results is required, because factors such as specimen type, transport conditions, the presence of commensal organisms, and co-detection of multiple pathogens may influence diagnostic accuracy. Particularly, for bacterial pathogens, the detection of nucleic acids alone does not necessarily indicate active infection (17, 18).
Despite these considerations, studies simultaneously analyzing B. pertussis, other major bacterial respiratory pathogens, and atypical pneumonia–causing bacteria in children and adolescents at a regional level in Korea remain scarce. Therefore, in this study, we conducted molecular detection–based analyses targeting seven pathogens, including six major bacterial respiratory pathogens (B. pertussis, M. pneumoniae, Chlamydia pneumoniae, S. pneumoniae, Legionella pneumophila, and H. influenzae) and the pertussis-related species B. holmesii, using respiratory specimens collected from children and adolescents in Busan. By applying molecular diagnostic methods, we aimed to characterize pathogen distribution patterns, age group–specific detection profiles, and co-detection patterns among bacterial respiratory pathogens. These analyses are intended to elucidate the epidemiologic characteristics of bacterial respiratory infections accompanying the pertussis resurgence and provide baseline data to inform future multi-pathogen–based diagnostic and treatment strategies.
MATERIALS AND METHODS
Study population and specimens
In total, 607 respiratory specimens were obtained from patients aged < 19 years who were diagnosed as pertussis-positive at private medical institutions, including clinics and hospitals, in Busan, Korea, between July and December 2024. The specimens comprised 602 nasopharyngeal swabs and five sputum samples. All specimens were residual samples remaining after routine diagnostic testing and were transported to the laboratory under refrigerated conditions via a specimen transport service.
Most nasopharyngeal swab specimens were submitted in viral transport medium (VTM). Upon arrival, specimens were processed immediately or stored at −75°C until further analysis.
At the participating private medical institutions, pertussis diagnosis was performed using PCR-based molecular assays according to the diagnostic protocols of each laboratory; some institutions used PCR assays targeting the insertion sequence IS481. In this study, the submitted residual specimens were further analyzed using PCR targeting the pertussis toxin gene (ptxA) of B. pertussis. Therefore, discrepancies between the initial pertussis diagnosis at the medical institutions and the results obtained in this study may be attributed to differences in diagnostic targets, assay methods, specimen storage and transport conditions, and timing of testing.
Target pathogens
To comprehensively evaluate the major causative agents of bacterial respiratory infections, molecular testing was performed for seven pathogens, including six bacterial respiratory pathogens and one pertussis-related species. The target pathogens included B. pertussis, M. pneumoniae, C. pneumoniae, L. pneumophila, S. pneumoniae, and H. influenzae. In addition, B. holmesii was included to enable differentiation of pertussis-like infections caused by closely related species.
These pathogens were selected because they are established causes of respiratory infections or pertussis-like clinical symptoms in children and adolescents, and their detection may reveal co-detection patterns not captured by routine pertussis diagnostic algorithms.
Specimen processing and nucleic acid extraction
Nasopharyngeal swab specimens submitted in VTM were directly subjected to nucleic acid extraction by aliquoting a defined volume of the transport medium. Because sputum specimens are highly viscous and unsuitable for direct extraction, they were diluted and homogenized using sterile phosphate-buffered saline, after which the clarified supernatant was collected for analysis.
Nucleic acids were extracted from pretreated specimens using an automated extraction system (Nextractor NX-48; Genolution, Korea) with the NX-48 Bacterial DNA Kit, according to the manufacturer’s instructions. Extracted nucleic acids were either used immediately for molecular analysis or stored at −75°C until testing.
Detection of bacterial respiratory pathogens
Extracted nucleic acids were used as templates for the detection of bacterial respiratory pathogens. Six major bacterial respiratory pathogens were analyzed using real-time PCR assays with the PowerChek™ Bacterial Pneumonia Real-time PCR Kits I and II (KogeneBiotech, Korea). These assays are designed for the simultaneous detection of major respiratory bacterial pathogens, including B. pertussis (ptxA), M. pneumoniae, C. pneumoniae, L. pneumophila, S. pneumoniae, and H. influenzae. Each reaction was prepared by adding 5 μL of extracted nucleic acid and 5 μL of primer–probe mix to the premix, in accordance with the manufacturer’s protocol.
PCR amplification was conducted under the following cycling conditions: 50°C for 2 min (1 cycle), 95°C for 10 min (1 cycle), 95°C for 15 sec and 60°C for 1 min(40 cycles). Amplification was performed using the ABI 7500 Fast Real-Time PCR System (Applied Biosystems, USA). Pathogen detection was determined based on amplification curves and cycle threshold (Ct) values, with positivity criteria defined according to the manufacturer’s instructions.
B. holmesii was analyzed using conventional PCR targeting hls1001, in accordance with the standard operating procedures of the Korea Disease Control and Prevention Agency. PCR products were confirmed by agarose gel electrophoresis. Co-detection was defined as the presence of two or more pathogens in a single specimen.
RESULTS
Detection of bacterial respiratory pathogens according to specimen type
Among the 607 respiratory specimens analyzed, 602 (99.2%) were nasopharyngeal swabs and five (0.8%) were sputum specimens. At least one bacterial respiratory pathogen was detected in 366 specimens, corresponding to an overall detection rate of 60.3%.
By specimen type, bacterial pathogens were detected in 361 of 602 nasopharyngeal swab specimens, yielding a detection rate of 60.0%. In contrast, all five sputum specimens tested positive for at least one bacterial respiratory pathogen, corresponding to a detection rate of 100.0%; however, the number of sputum specimens was limited.
In nasopharyngeal swab specimens, B. pertussis showed the highest detection rate at 33.2% (200/602), followed by H. influenzae at 19.1% (115/602), B. holmesii at 12.8% (77/602), S. pneumoniae at 12.1% (73/602), and M. pneumoniae at 8.5% (51/602). C. pneumoniae and L. pneumophila were detected at low frequencies of 0.8% (5/602) and 0.2% (1/602), respectively.
Although sputum specimens accounted for only five cases, at least one bacterial respiratory pathogen was detected in all samples.
Age-group–specific detection rates of bacterial respiratory pathogens
Of the 607 respiratory specimens analyzed, 366 were positive for at least one bacterial respiratory pathogen, yielding an overall detection rate of 60.3%. Study participants were categorized into four age groups: preschool children (0–7 years), elementary school students (8–13 years), middle school students (14–16 years), and high school students (17–19 years).
The highest detection rate was observed in the preschool group, with 20 of 29 specimens testing positive (69.0%); however, the sample size in this group was limited. In the elementary school group, 256 of 430 specimens were positive, corresponding to a detection rate of 59.5%. Moreover, this group accounted for the largest proportion of positive cases. The detection rate in the middle and high school groups were 63.3% (69/109) and 53.8% (21/39), respectively.
Although detection rates were generally comparable across age groups, the absolute number of positive cases was highest among elementary school students, indicating a concentration of bacterial respiratory infections in school-aged children and adolescents.
Detection rates and distribution of bacterial respiratory pathogens
Among the 607 respiratory specimens analyzed for six major bacterial respiratory pathogens and one pertussis-related species, marked differences were observed in pathogen-specific detection rates and distribution patterns (Table 1, Fig. 1). B. pertussis was the most frequently detected pathogen, identified in 205 specimens, and remained the predominant pathogen across all age groups. This was followed by H. influenzae (119 cases), B. holmesii (79 cases), S. pneumoniae (73 cases), and M. pneumoniae (55 cases). In contrast, C. pneumoniae and L. pneumophila were detected at very low frequencies, with 5 and 1 cases, respectively.
Table 1.
Distribution of detected bacterial respiratory pathogens by age group
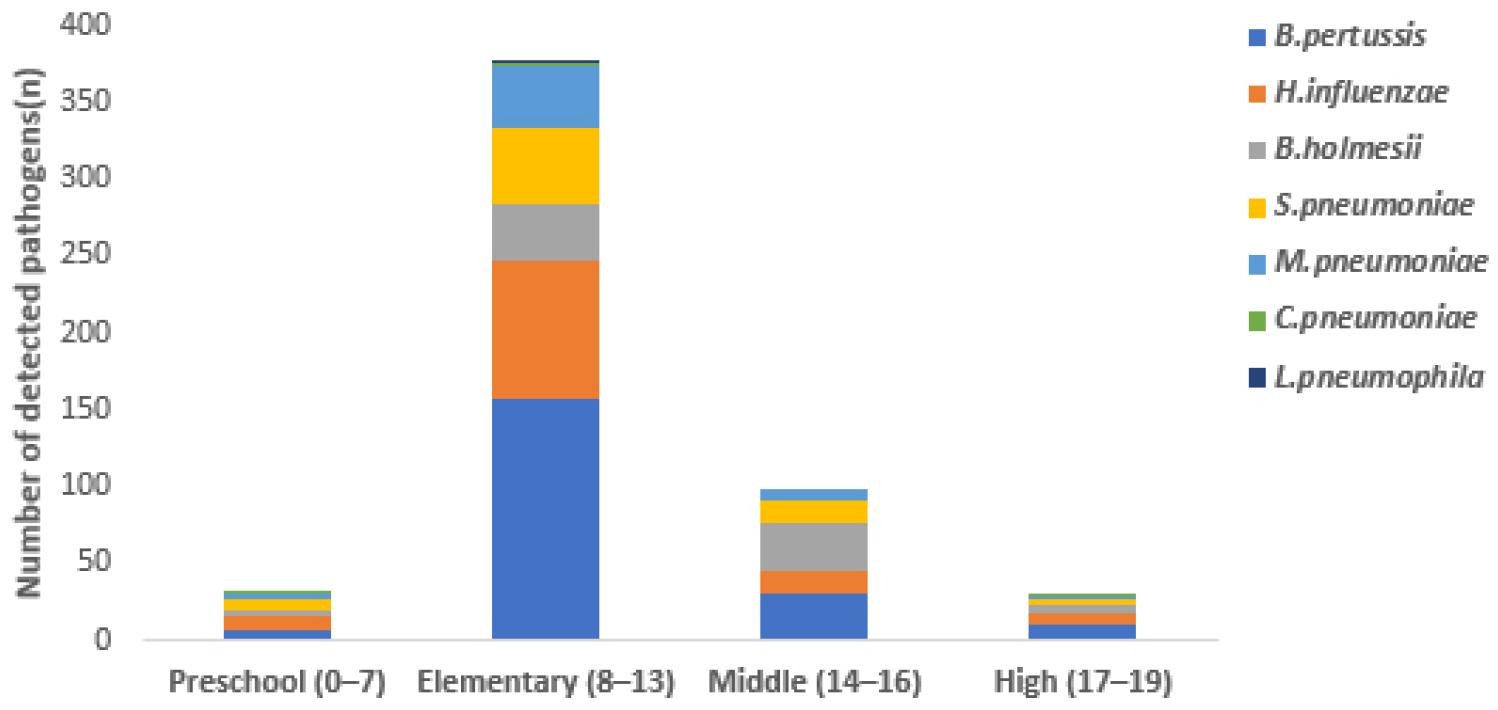
Fig. 1
Age group–specific distribution of detected bacterial respiratory pathogens.
Bordetella pertussis (B. pertussis), Haemophilus influenzae (H. influenzae), Bordetella holmesii (B. holmesii), Streptococcus pneumoniae (S. pneumoniae), Mycoplasma pneumoniae (M. pneumoniae), Chlamydia pneumoniae (C. pneumoniae), Legionella pneumophila (L. pneumophila).
Age group–specific analysis revealed that most bacterial pathogens were more frequently detected in the elementary and middle school age groups (Table 1). In particular, B. pertussis and H. influenzae were most prevalent among elementary school-aged children, whereas B. holmesii accounted for a relatively higher proportion of detections in the middle school age group. These age-specific distribution patterns are illustrated in Fig. 1. C. pneumoniae and L. pneumophila were detected sporadically across all age groups, without a clear age-specific trend.
Patterns of bacterial co-detection
Patterns of bacterial co-detection were analyzed among the 366 specimens in which at least one bacterial respiratory pathogen was detected. Single-pathogen detection was identified in 224 cases (61.2%), whereas co-detection involving two or more bacterial pathogens was observed in 142 cases (38.8%) (Table 2).
Table 2.
Patterns of single detection and bacterial co-detection
Among the co-detection patterns, dual-pathogen co-detections were the most common, accounting for 115 cases (31.4% of all positive specimens). The most frequent dual-pathogen combination was B. pertussis and H. influenzae (41 cases), followed by B. pertussis and S. pneumoniae (16 cases), M. pneumoniae and H. influenzae (10 cases), and S. pneumoniae and H. influenzae (8 cases) (Table 2).
Triple-pathogen co-detections were identified in 25 cases (6.8%), with the combination of B. pertussis, S. pneumoniae, and H. influenzae being the most frequent (9 cases). In addition, several triple-pathogen combinations involving M. pneumoniae were observed (Table 2). Quadruple-pathogen co-detections were rare, occurring in only two cases (0.5%), both involving combinations that included H. influenzae and B. holmesii (Table 2).
Overall, these findings demonstrate that, in pertussis-positive specimens, not only single-pathogen infections but also diverse patterns of bacterial co-detection are frequently observed, with multi-pathogen co-detection centered on B. pertussis being particularly prominent (Table 2).
DISCUSSION
This study analyzed the distribution and co-detection patterns of major bacterial respiratory pathogens in pertussis-positive respiratory specimens obtained from children and adolescents in Busan, Korea, during 2024, providing a descriptive overview of bacterial respiratory infections accompanying the recent pertussis resurgence.
Most specimens analyzed in this study were nasopharyngeal swabs, consistent with previous reports indicating that nasopharyngeal specimens are the standard sample type for respiratory infection surveillance in pediatric and adolescent populations (1, 17). During pertussis outbreaks, nasopharyngeal swabs provide high sensitivity for the detection of B. pertussis(6, 9), and a similar pattern was observed in the present study. However, the small number of sputum specimens limited comparisons of diagnostic performance according to specimen type.
Age group–specific analysis indicated that the detection of bacterial respiratory pathogens was concentrated in school-aged children and adolescents. However, because this study was restricted to pertussis-positive specimens, this distribution likely reflects the characteristics of the study population rather than true epidemiologic differences. Therefore, age-specific findings should be interpreted with caution. Given the study design, the high detection rate of B. pertussis was expected; therefore, this study focused on the distribution and co-detection patterns of other bacterial respiratory pathogens accompanying pertussis. The detection of B. holmesii, a closely related species, suggests that pertussis-like illness cannot always be attributed to a single pathogen. In this study, B. holmesii was detected in 79 cases, representing a relatively high frequency. B. holmesii has been reported as a causative agent of pertussis-like illness in several countries and can present with clinical features similar to those of B. pertussis(19). Importantly, B. holmesii shares the insertion sequence IS481, a commonly used molecular target for pertussis diagnosis. Consequently, in the absence of species-specific assays, infections caused by B. holmesii may be misclassified as B. pertussis(9, 19). Because IS481-targeted PCR assays are not species-specific and can detect both organisms, differentiation may not occur during routine diagnostic testing. Previous studies have suggested that a substantial proportion of patients with pertussis-like illness are infected with B. holmesii(19). In the present study, additional molecular analysis of specimens initially identified as pertussis-positive confirmed B. pertussis through detection of ptxA. Therefore, the observed co-detection of B. pertussis (ptxA-positive) and B. holmesii is unlikely to be attributable to cross-reactivity and may instead reflect the true co-presence of both organisms that were not differentiated at the initial stage of diagnosis.
These findings suggest that surveillance systems based solely on IS481 detection may overestimate the true burden of B. pertussis while underrepresenting infections caused by other Bordetella species. In addition, B. holmesii is not directly targeted by currently available pertussis vaccines, which represents an important distinction from B. pertussis(20). Collectively, these findings highlight the need to consider B. holmesii in the interpretation of pertussis surveillance data and in the development of diagnostic strategies. In contrast, H. influenzae and S. pneumoniae were detected at lower frequencies than those reported in previous studies of nasopharyngeal colonization (15, 16). This discrepancy may be attributable to the predominance of upper respiratory specimens and the use of VTM in this study. Antibiotic components in VTM may reduce bacterial viability and recovery, potentially affecting the molecular detection of bacterial pathogens (18).
Therefore, the findings of this study should be interpreted as complementary to, rather than directly comparable with, the initial diagnostic results obtained at private medical institutions.
The primary objective of this study was to characterize pathogen co-detection patterns in respiratory infections and provide baseline data to inform future treatment strategies. Pertussis is generally responsive to macrolide therapy (6, 9), whereas M. pneumoniae has shown a sustained increase in macrolide-resistant strains in Korea and elsewhere, contributing to treatment failure and prolonged disease courses (10, 11, 12, 13, 14). In cases of co-detection involving B. pertussis and M. pneumoniae, clinical outcomes may vary depending on the pathogen predominantly driving the infection and guiding treatment decisions.
Consistent with this concern, co-detection of B. pertussis with M. pneumoniae, H. influenzae, and S. pneumoniae was frequently observed in our study. These findings indicate that diagnostic and therapeutic strategies based solely on a single detected pathogen do not adequately reflect the complexity of respiratory infections in pediatric patients. Accordingly, our results support the use of multiplex PCR-based diagnostic approaches capable of simultaneous detection of multiple pathogens (17).
Multiplex PCR assays enable the simultaneous detection of multiple respiratory pathogens and are now widely used in clinical diagnostics and surveillance studies (17). This approach provides high analytical sensitivity and facilitates rapid identification of multiple pathogens within a single specimen, which is particularly useful for characterizing co-detection patterns. However, nucleic acid–based detection does not necessarily indicate active infection, because some detected microorganisms may represent asymptomatic carriage or colonization of the upper respiratory tract rather than true etiologic pathogens (15, 16). Therefore, interpretation of multiplex PCR results should be undertaken cautiously, in conjunction with clinical and epidemiologic information. In this study, although multiplex PCR allowed comprehensive detection of bacterial respiratory pathogens, it was not possible to distinguish between true infection and colonization based solely on molecular findings.
This study has some limitations. As a retrospective analysis, it was constrained by limited information on clinical symptoms, antibiotic treatment history, and antimicrobial resistance profiles. Nevertheless, by systematically describing the distribution and co-detection patterns of bacterial respiratory pathogens among children and adolescents during a pertussis outbreak, our findings provide baseline data that may inform the development of diagnostic strategies and treatment guidelines that account for potential antimicrobial resistance.
CONCLUSION
This study analyzed the distribution and co-detection patterns of major bacterial respiratory pathogens in pertussis-positive respiratory specimens obtained from children and adolescents in Busan, Korea, during 2024. Among the analyzed pertussis-positive respiratory specimens, B. pertussis was detected across all age groups, with a higher proportion observed among school-aged children and adolescents. Co-detection of two or more bacterial pathogens occurred in a substantial proportion of specimens, highlighting the limitations of diagnostic and therapeutic approaches focused on a single pathogen. In particular, co-detection involving M. pneumoniae underscores the importance of considering antimicrobial resistance when selecting treatment strategies. Taken together, these findings provide baseline data to support enhanced surveillance of bacterial respiratory infections and the development of diagnostic and therapeutic strategies based on multi-pathogen detection.




