INTRODUCTION
Klebsiella pneumoniae (Kp) is an opportunistic Gram-negative pathogen that causes a broad spectrum of infections, including pneumonia, urinary tract infections, liver abscess, and sepsis, and represents a major global health threat due to its increasing prevalence of multidrug resistance (1). In particular, the emergence of hvKp has drawn considerable attention. Unlike classical Kp (cKp), hvKp strains possess enhanced pathogenicity and can cause severe, community-acquired infections even in young and otherwise healthy individuals (2). HvKp is characterized by a distinctive combination of genetic and phenotypic traits. Genetically, hvKp typically carries a set of virulence plasmid–encoded determinants, including rmpA/rmpA2, which upregulate capsular polysaccharide (CPS) synthesis, and siderophore systems such as aerobactin (iucABCD/iutA) and salmochelin (iroBCDN)—recognized as among the most reliable molecular markers of hvKp. Additional virulence-associated loci, including peg-344, kfu, and allS, are also enriched in hvKp and contribute to its invasive phenotype (3). Phenotypically, hvKp is characterized by hypermucoviscosity, demonstrable by the string test, and by overproduction of CPSs, which together confer immune evasion, resistance to complement-mediated killing, and increased potential for metastatic infection (4, 5). These genetic signatures and CPS-driven mucoviscous phenotype collectively serve as the standardized framework defining hvKp and distinguishing it from classical strains.
Phenotypic plasticity—especially variation in capsule expression—is frequently observed among Enterobacteriaceae. For instance, in a murine gut-colonization model, spontaneous capsule loss in Kp occurred rapidly, with differences depending on the pathotype (6). Such phenotypic switching can be associated with altered virulence traits. Moreover, capsule expression has been shown to enhance bacterial survival under immune pressures: in Escherichia coli carrying the K1 capsule, serum-mediated killing was reduced, and removal of the capsule sensitized bacteria to immune responses (7). Importantly, bacterial capsules not only protect against host defenses but also drive adaptation to novel environments. Given that capsule-dependent adaptation can modify multiple virulence-associated phenotypes in structured environments (7), the observed capsule loss likely results in diminished exopolysaccharide production. This reduction in exopolysaccharide is known to facilitate biofilm formation by increasing surface adhesion, providing a coherent mechanistic explanation for the enhanced biofilm phenotype.Additionally, mutations in the wzc gene have been identified in clinical hvKp isolates as drivers of reduced CPS synthesis and altered mucoid phenotype, serving as a potential host-adaptive strategy (8). However, the broader implications of capsule heterogeneity in hvKp for antibiotic resistance and macrophage interactions remain insufficiently understood.
In this study, we describe the emergence of irregular, small-sized hvKp colonies with reduced mucoviscosity. We examined their virulence gene expression, susceptibility to polymyxin B, and functional behavior in macrophage infection models. Our findings demonstrate that despite showing no significant differences in canonical virulence gene expression, these phenotypically altered colonies exhibited markedly increased resistance to polymyxin B and significantly enhanced phagocytosis and intracellular survival. These results highlight the complex and dynamic nature of hvKp phenotypic variation and suggest that capsule-deficient variants may represent a previously underappreciated survival strategy within the host.
MATERIALS AND METHODS
Cell culture
The murine macrophage cell line RAW264.7 (TIB-71; American Type Culture Collection) was used in this study. Cells were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM; Welgene, Gyeongsan, South Korea), supplemented with 10% fetal bovine serum (FBS; Welgene) and 1% penicillin/streptomycin (Gibco BRL, Grand Island, NY, USA). Cells were incubated at 37°C in a humidified atmosphere containing 5% CO2.
Bacterial strains and culture
An hvKp strain was isolated from blood specimens of patients diagnosed with liver abscesses at Keimyung University Dongsan Hospital (Daegu, South Korea). Bacterial identification and antimicrobial susceptibility testing were performed using the VITEK 2 automated system (bioMérieux, Lyon, France). Production of extended-spectrum β-lactamase (ESBL) was confirmed via the agar dilution method using cefotaxime, ceftazidime, and ceftazidime in combination with clavulanate, following Clinical and Laboratory Standards Institute (CLSI) guidelines. Capsular serotypes and virulence-associated genes were determined by PCR. Hypervirulent characteristics were assessed via the string test and PCR amplification of rmpA, magA, and aerobactin genes. Based on serotyping results, hvKp belonged to the K1 serotype. Bacterial strains were cultured in Luria-Bertani (LB) broth (244620, Difco, Rockville, MD, USA) for liquid cultures and on LB agar (244520, Difco) as solid medium at 37°C. For long-term storage, bacterial stocks were prepared in LB broth supplemented with 15% (v/v) glycerol (Sigma-Aldrich, St. Louis, MO, USA) and stored at –80°C. For infection experiments, bacteria were first cultured in LB broth for 16–18 h at 37°C. The overnight cultures were then diluted in fresh LB broth and grown to an optical density at 600 nm (OD600) of 0.1. The resulting bacterial suspensions were aliquoted, supplemented with 15% glycerol, and stored at –80°C. One aliquot was thawed and colony-forming units (CFUs) were quantified prior to use in infection assays.
String test
Bacterial strains were cultured in LB broth at 37°C with shaking at 220 rpm for 18 h. Bacteria cultures were centrifuged at 10,000 rpm for 3 min, and the supernatant was removed. The bacterial pellet was resuspended and adjusted to an OD600 of 0.1 using phosphate buffered saline (PBS). The suspension was serially diluted tenfold in PBS and 100 µL of each dilution was spread onto LB agar plates and MacConkey agar (211387, Difco) plates. After overnight incubation at 37°C, single colonies were picked with a sterile inoculation loop. The loop was lifted vertically to measure the length of the viscous string formed; a string length ≥ 5 mm was considered positive.
Bacterial growth curve
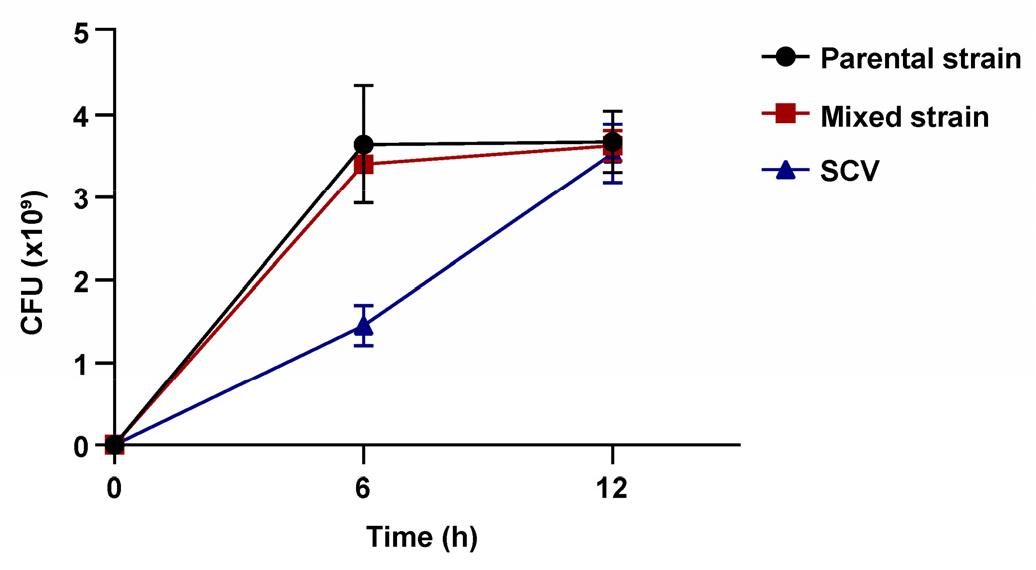
Bacterial strains were cultured in LB broth at 37°C with shaking at 220 rpm for 18 h. The cultures were adjusted to an OD600 of 0.1 using LB broth. The suspension was then diluted 1:50 in LB broth and incubated at 37°C with shaking. CFUs were measured at 0, 6, and 12 h.
Intracellular CFU assay
RAW264.7 cells were infected with hvKp at a multiplicity of infection (MOI) of 40 for 1 h at 37°C in 5% CO2 using antibiotic-free medium. The cells were then washed three times with PBS and incubated for an additional 30 min in medium containing 100 μg/mL gentamicin (15710-072, Gibco). After washing, the cells were lysed with 1% saponin in distilled water. Ten-fold serial dilutions of the lysates were plated on LB agar and incubated overnight at 37°C to determine CFUs. For phenotypic classification experiments, macrophages were infected with the mixed strain, and intracellular bacteria were recovered 6 h post-infection. Colonies were classified based on colony morphology as parental-type or SCVs. Colonies were counted after 24 h of incubation.
Mucoviscosity assay
Mucoviscosity of bacteria was assessed by measuring the OD600 of the supernatant after low-speed centrifugation of bacteria resuspended in PBS (9). Cultures were grown in LB broth for 16–18 h at 37°C with shaking at 220 rpm and diluted 1:100 in fresh LB broth. The cells were harvested by centrifugation at 10,000 rpm for 3 min and resuspended in PBS to an OD600 of 1.0. The suspensions were then centrifuged at 1,000 g for 5 min, and the OD600 of the supernatant was measured. Mucoviscosity was quantified as the ratio of the OD600 of the suspension to the initial OD600.
Minimum inhibitory concentration (MIC) assay
MIC assays were performed to evaluate bacterial susceptibility to gentamicin, ampicillin (MB-A4128, MB Cell), colistin sulfate (MB-C6212, MB cell), meropenem (PHR1772, Sigma-Aldrich), and polymyxin B (81271, Sigma-Aldrich). Bacterial isolates were initially cultured in LB broth for 16–18 h at 37°C in a shaking incubator at 220 rpm. Overnight cultures were subcultured in fresh broth and grown to the early–mid logarithmic phase. Bacterial cells were directly adjusted to the OD600 of 0.1 (approximately 0.5 McFarland standard) using cation-adjusted Mueller–Hinton broth (CA-MHB; 90922, Sigma- Aldrich). The bacterial suspension was diluted to achieve a final inoculum of approximately 5 × 10⁵ CFU/mL. Serial two-fold dilutions of each antibiotic were prepared in CA-MHB and mixed with the bacterial inoculum. MIC assays were incubated at 37°C for 16–18 h under static conditions. MIC values were determined by visual inspection, defined as the lowest antibiotic concentration that completely inhibited visible bacterial growth. OD600 was subsequently measured using a Synergy/HTX multimode microplate reader (BioTek Instruments, Inc., Winooski, VT, USA) as a supplementary assessment.
Preparation of genomic DNA
For genomic DNA preparation, we employed a heat treatment–based method (10) with minor modifications. A single colony was inoculated into 500 µL of LB broth in 1.5 mL microtubes and incubated overnight at 37°C in an incubator (MIR-253, Sanyo, Japan) equipped with a rotator (CRT-350, Jeiotech, Korea) at 30 rpm. The cultures were centrifuged at 13,000 rpm for 3 min using a centrifuge (5424, Eppendorf, Germany). The supernatant was carefully aspirated in a biosafety cabinet (JSCB-1200SB, JSR, Korea), and the pellet was resuspended in 100 µL of nuclease-free water (129114, Qiagen, Hilden, Germany). The suspension was incubated at 100°C for 10 min in a heating block (ALB64, FINEPCR, Korea), followed by centrifugation at 13,000 rpm for 1 min. An aliquot of 80 µL of the supernatant was transferred to a new 1.5 mL tube and either stored at −20°C or used immediately for PCR.
Multiplex PCR of bacterial virulence factors
Primers and detailed reaction conditions are provided in Supplementary Tables 1 and 2. (3, 11, 12, 13, 14). All reactions were prepared in 0.2 mL PCR tubes. Multiplex PCR assays were performed in two groups based on annealing temperature: at 50°C for magA, allS, iucA, K2, rmpA, and ybtS, and at 56°C for iutA, kfu, iroN, entB, mrkD, and rmpA2. DNA polymerase, 5× Green Buffer, MgCl₂ (M8295, Promega), and dNTP mix (U151B, Promega) were all obtained from Promega (Madison, WI, USA). Agarose gels were prepared by dissolving 0.75 g agarose (REF. V3125, Promega) in 50 mL of 1× TAE buffer (diluted from 50× TAE, Cat. No. T2002, Biosesang, Seongnam, Korea) using a microwave, followed by addition of 5 µL EcoDye™ (ES301-1000, Biofact, Daejeon, Korea). The mixture was poured into a gel tray with combs and allowed to solidify for at least 40 min. Gels were placed in a Mupid-2 Plus electrophoresis system (Advance, Tokyo, Japan) containing 1× TAE buffer. PCR products (10 µL out of the 20 µL total reaction volume) and a DNA ladder (D-1030, Bioneer, Daejeon, Korea) were loaded. Electrophoresis was conducted at 100 V for 40 min, and gels were visualized using a multifunctional imager (SH-523, SHST, China) and program (SHST capture) under UV plate mode with a 20 ms exposure time.
Statistical analysis
Data are presented as mean ± standard deviation and were obtained from three independent experiments. The Student’s t-test or analysis of variance was used to analyze the data. Statistical analyses were performed, and graphs were made using GraphPad Prism 7.0 software for Windows (GraphPad Software Inc., La Jolla, CA, USA). P-values <0.05, 0.01, and 0.001 were considered statistically significant.
RESULTS
Single hvKp strain generates distinct phenotypic variants
We infected RAW264.7 cells with a single hvKp strain and identified the following three distinct colony variants: (1) normal-sized, mucoid colonies; (2) normal-sized, nonmucoid colonies; and (3) small-sized, nonmucoid colonies. Each variant was isolated as a single colony and subsequently used to infect RAW264.7 cells. After cell lysis, normal-sized mucoid colonies retained their original phenotype (parental strain), whereas normal-sized nonmucoid colonies (mixed strain) displayed heterogeneous colonies. In contrast, small nonmucoid colonies, corresponding to SCVs, consistently retained their small, nonmucoid phenotype. In addition, string test analysis revealed that the parental strain exhibited the highest degree of hypermucoviscosity, followed by the mixed strain, whereas SCVs remained nonviscous (Fig. 1). Although the SCV showed delayed growth at 6 h, the growth curves of all three strains converged to similar levels by 12 h (Fig. 2). Taken together, these results indicate that a single hvKp strain can produce diverse phenotypic variants.
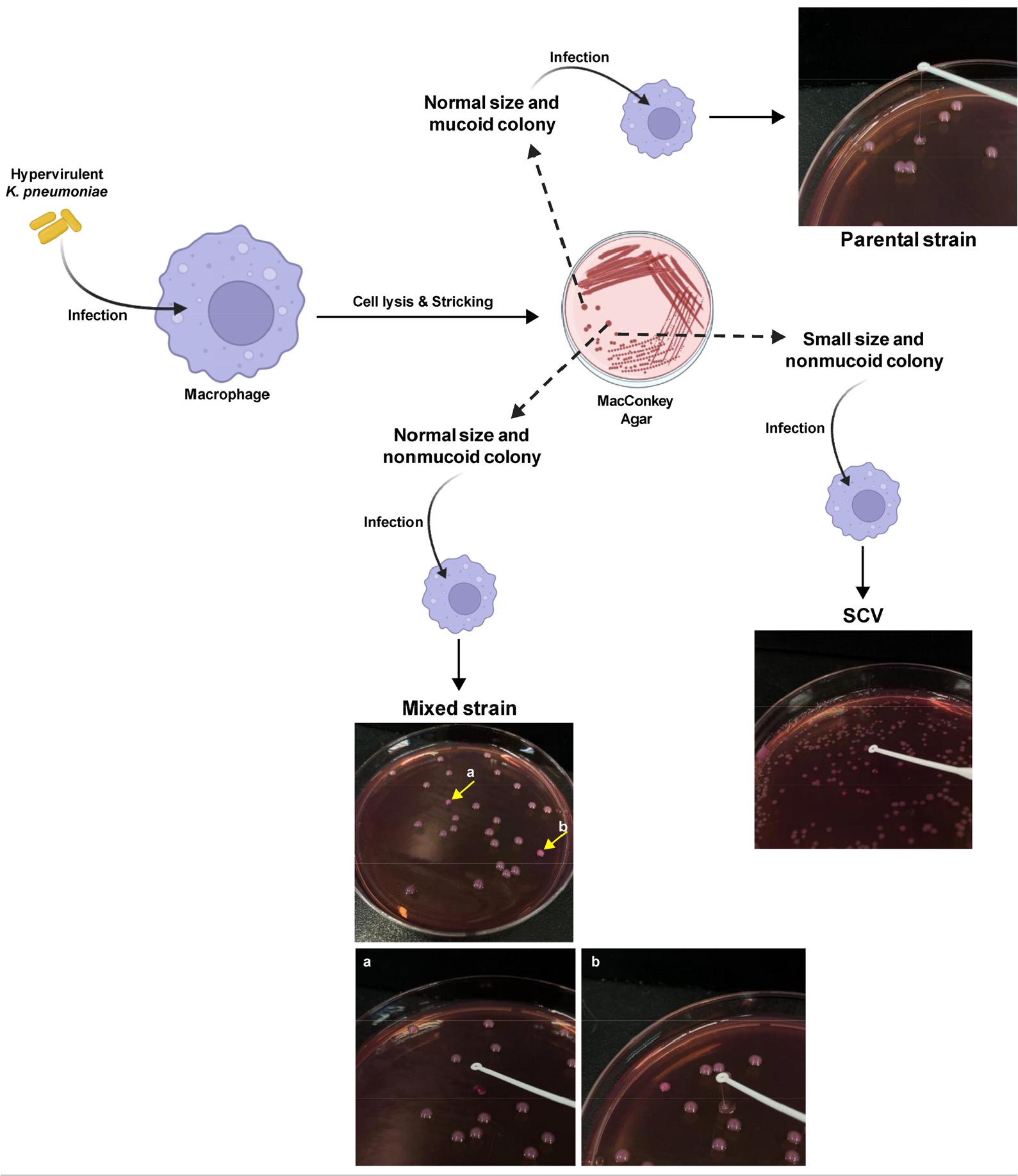
Fig. 1
Morphological characterization of variants. RAW264.7 cells were infected with hvKp (MOI 40) for 1 h, after which the cells were lysed and plated on MacConkey agar. Three distinct colony morphotypes were observed: (i) normal-sized, mucoid (parental) colonies; (ii) normal-sized, nonmucoid colonies; and (iii) small-sized, nonmucoid colonies corresponding to SCVs. Each morphotype was isolated as a single colony and used to reinfect macrophages to assess phenotypic stability. The string test was performed on each variant using a sterile loop—panel (a) shows a negative string test, whereas panel (b) shows a positive result. Created with BioRender.com.
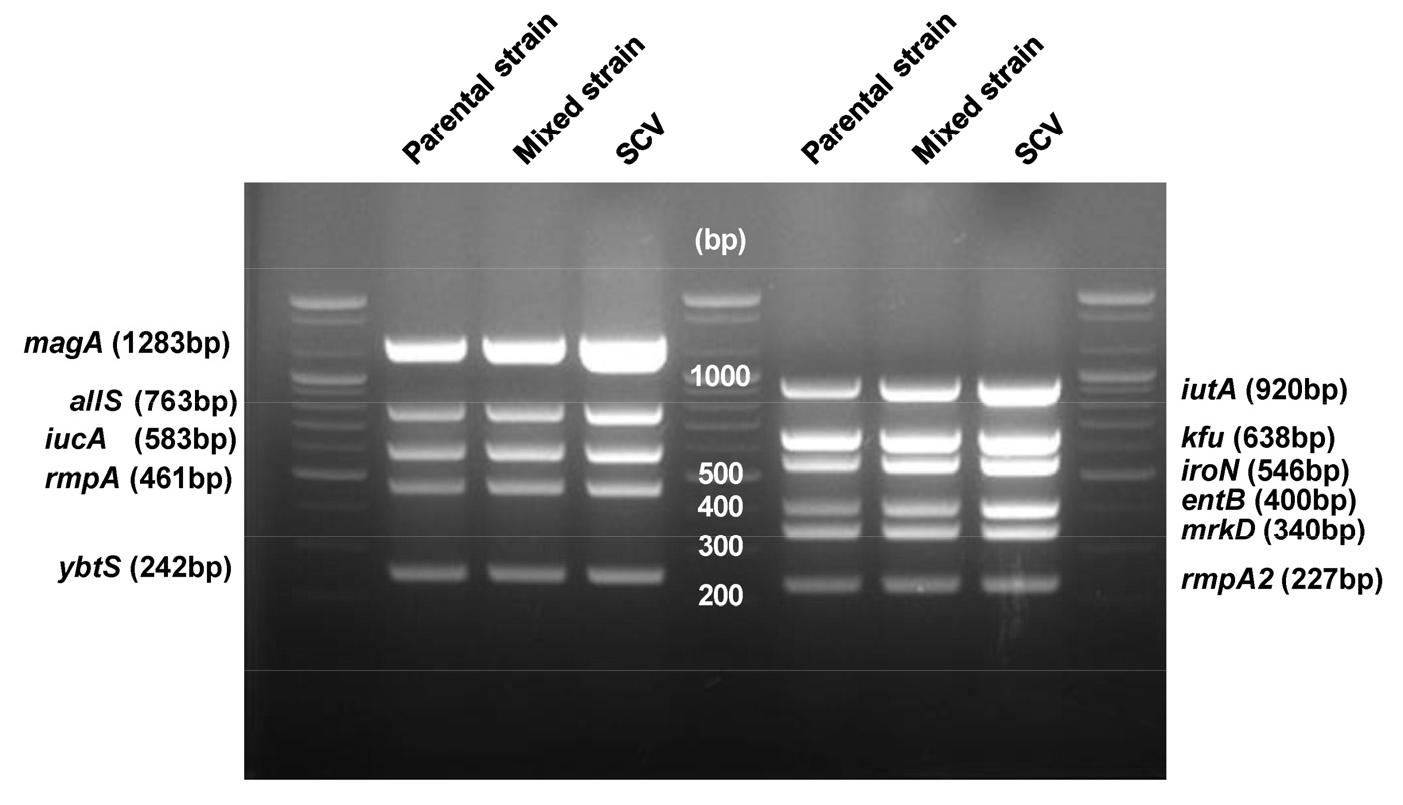
Multiplex PCR reveals identical virulence determinants in variants
Multiplex PCR confirmed the presence of multiple virulence-associated genes in the Kp isolates. At an annealing temperature of 50°C, amplification products corresponding to magA (1283 bp), allS (764 bp), iucA (583 bp), rmpA (461 bp), and ybtS (242 bp) were detected. Reactions performed at 56°C yielded amplifications of iutA (920 bp), kfu (638 bp), iroN (546 bp), entB (400 bp), mrkD (340 bp), and rmpA2 (227 bp). Notably, all three isolates exhibited identical virulence gene profiles, with no significant variations among the 11 genes tested (Fig. 3). These findings indicate that the Kp isolates retain a conserved set of virulence determinants, suggesting a stable genetic background for key pathogenic traits across variants.
Differential mucoviscosity phenotypes among hvKp variants
When the three hvKp variants were cultured in LB medium and subjected to centrifugation, the supernatants of the parental and mixed strains appeared turbid, whereas SCVs formed a visible pellet (Fig. 4A). To further confirm this observation, we performed a mucoviscosity assay, which demonstrated that SCVs exhibited the lowest mucoviscosity. For up to 6 h of incubation, the parental strain showed higher mucoviscosity than the mixed strain; however, after 9 h, both strains displayed comparable levels of mucoviscosity. Notably, the mucoviscosity of both strains decreased significantly after 18 h of incubation (Fig. 4B). These findings indicate that hvKp variants differ in their mucoviscosity phenotypes, with SCVs showing markedly reduced mucoviscosity and that the mucoviscosity of the parental and mixed strains dynamically declines over prolonged culture.
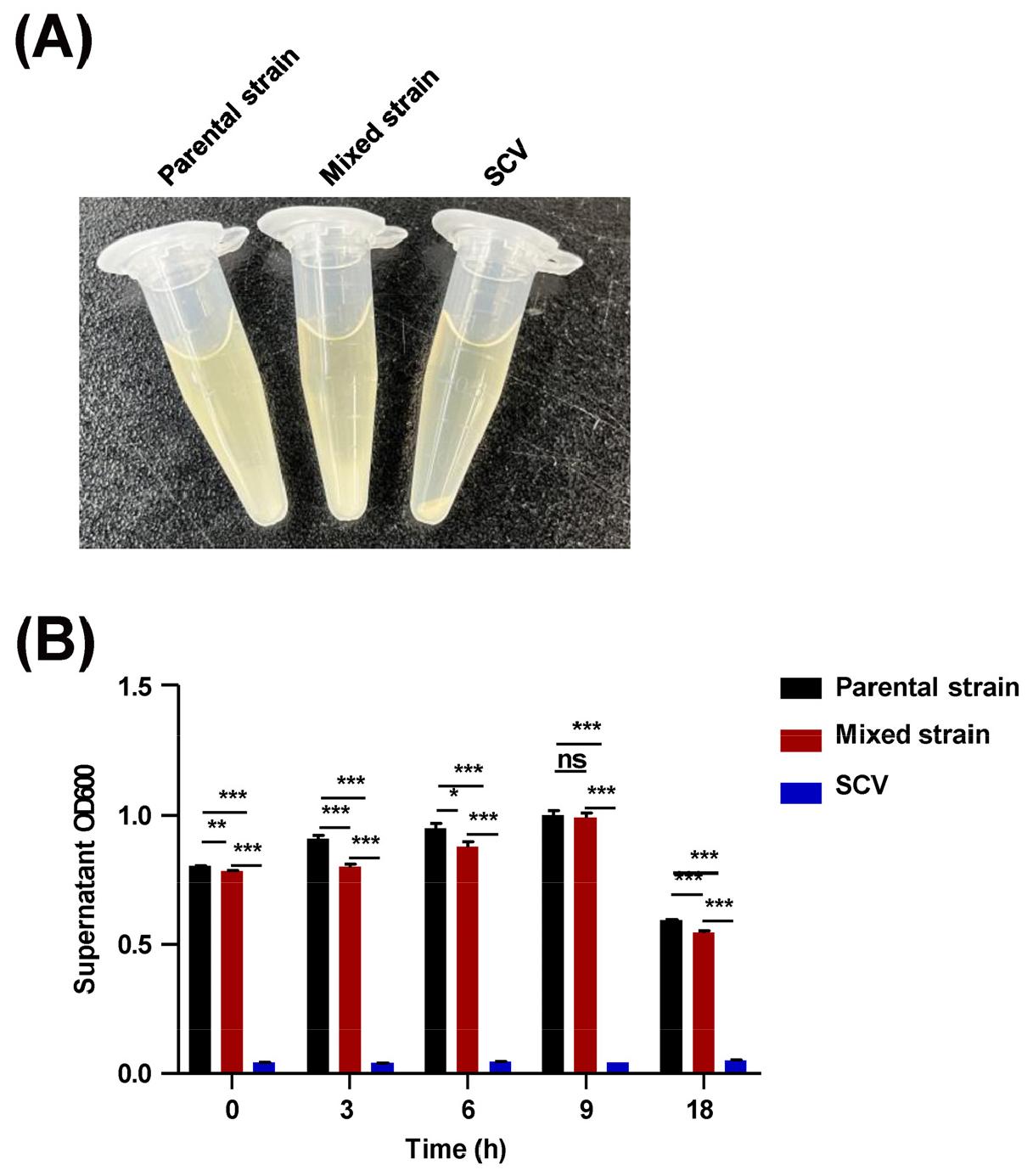
Fig. 4
Comparison of mucoviscosity among hvKp variants. (A) Image of bacterial cultures after centrifugation at 10,000 rpm for 3 min. (B) The mucoviscosity assay of three variants was measured at the indicated times. Data are expressed as the mean ± standard deviation of three independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001 according to analysis of variance. ns, not significant.
Altered phenotype is associated with differential antibiotic susceptibility
Recent studies suggest a potential association between phenotypic changes and antibiotic susceptibility (15). Therefore, we investigated whether the altered phenotypes observed after infection were related to antimicrobial resistance. In the MIC assay for polymyxin B, complete growth inhibition of the parental strain was observed at 5 μg/mL, the mixed strain and SCVs required 10 μg/mL for complete growth inhibition (Fig. 5A). All the strains were susceptible to gentamicin (Fig. 5B). In the ampicillin MIC assay, all strains were completely inhibited at 200 μg/mL, similar to the parental strain (Fig. 5C). In the colistin MIC assay, the parental strain and SCVs showed comparable susceptibility, with inhibition at 8 μg/mL, whereas the mixed strain displayed altered susceptibility, being inhibited at 4 μg/mL, potentially reflecting phenotypic variation within the mixed population (Fig. 5D). All the strains were susceptible to meropenem (Fig. 5E). These results indicate that the phenotypic alteration of the parental strain is associated with increased resistance to polymyxin B.
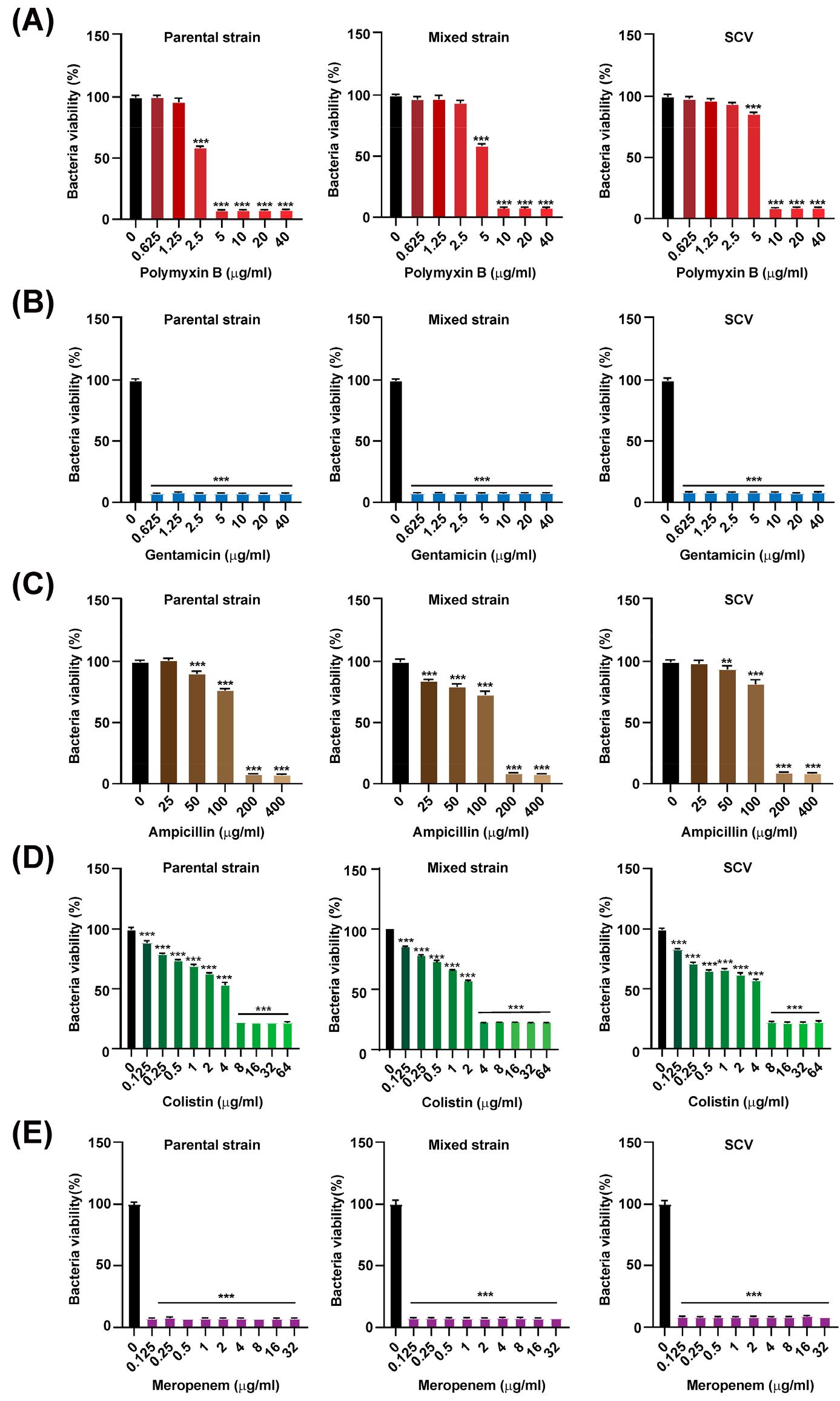
Fig. 5
Comparison of antimicrobial susceptibility among hvKp variants. (A) MIC of polymyxin B determined at concentrations ranging from 0.625 to 40 μg/mL. (B) MIC of gentamicin determined at concentrations ranging from 0.625 to 40 μg/mL. (C) MIC of ampicillin determined at concentrations ranging from 6.25 to 400 μg/mL. (D) MIC of colistin determined at concentrations ranging from 0.125 to 64 μg /mL. (E) MIC of meropenem determined at concentrations ranging from 0.125 to 32 μg/mL. Data are expressed as the mean ± standard deviation of three independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001 according to analysis of variance.
Differences in phagocytosis and intracellular survival among hvKp variants
To assess whether phenotypic variations influenced intracellular persistence, we measured intracellular bacterial survival at the time of phagocytosis (0 h) and at 6 and 24 h post-inoculation. Marked differences in bacterial uptake were evident at 0 h: the mixed strain showed the lowest uptake, the parental strain displayed intermediate uptake, and SCVs were internalized most efficiently (Fig. 6A). Following phagocytosis, intracellular bacterial survival was monitored for 6 h and 24 h. The mixed strain consistently displayed the lowest survival rate, the parental strain exhibited intermediate persistence, and SCVs maintained a markedly higher burden throughout the infection period. To quantitatively assess the robustness of phenotype switching, macrophages were infected with the mixed strain and intracellular bacteria were recovered for CFU enumeration and phenotypic classification. Notably, SCVs constituted the dominant subpopulation, accounting for 88.7% of the total recovered bacteria, whereas parental-type colonies represented only 11.3% (Fig. 6B). This SCV-dominant population distribution was consistently reproduced across independent experiments, supporting the robustness of SCV enrichment following macrophage infection. Taken together, these findings demonstrate that phenotypic switching in the parental strain regulates both phagocytic efficiency and intracellular persistence in a stepwise manner (mixed strain < parental strain < SCV), highlighting a critical trade-off in which mucoviscosity governs the balance between initial uptake and survival within host cells.
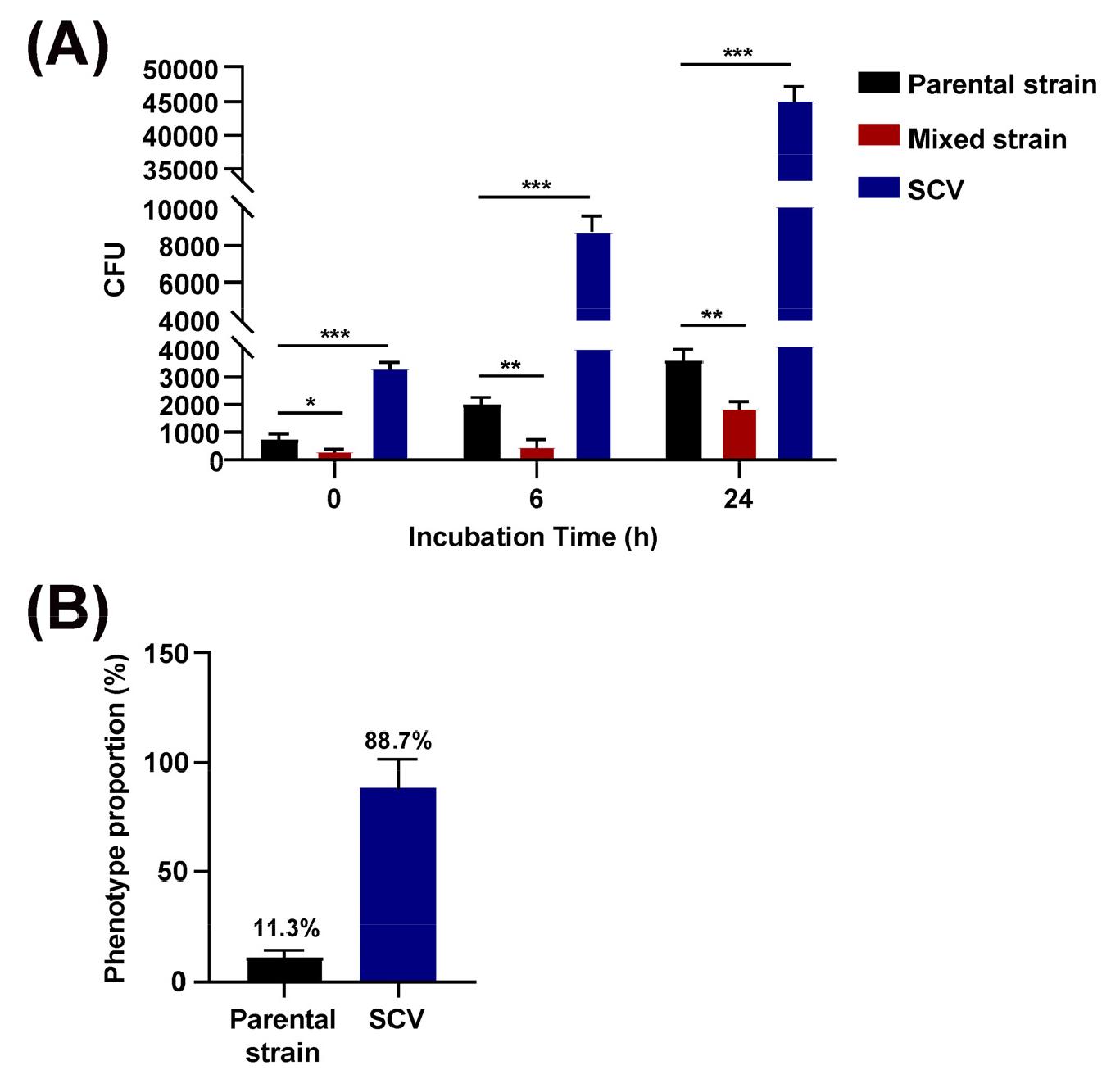
Fig. 6
Phenotypic switching influences phagocytosis efficiency and intracellular bacterial survival in macrophages. RAW264.7 cells were infected with each hvKp variant (MOI 40) for 1 h. (A) Intracellular bacterial survival was quantified by CFU enumeration at the time of phagocytosis (0 h), 6 h, and 24 h post-inoculation. (B) Following infection with the mixed strain, intracellular bacteria were recovered 6 h post-infection for CFU enumeration and phenotypic classification based on colony morphology. Data are expressed as the mean ± standard deviation of three independent experiments. Statistical significance was determined by one-way ANOVA followed by post hoc analysis. *P < 0.05, **P < 0.01, ***P < 0.001.
DISCUSSION
Our study demonstrated that hvKp undergoes phenotypic switching during macrophage infection, which profoundly affects colony morphology, mucoviscosity, antibiotic susceptibility, and intracellular persistence. We identified a stepwise relationship in which the mixed variant exhibited the lowest uptake and poorest survival, the parental strain showed intermediate phagocytosis and persistence, and SCVs were most efficiently phagocytosed while maintaining the highest intracellular burden. These results highlight the fundamental trade-off between extracellular immune evasion and intracellular adaptation.
Phenotypic switching enables bacterial adaptation during infection. The SCV is a slow-growing bacterial subpopulation characterized by atypical colony morphology, unusual biochemical traits, and reduced antibiotic susceptibility. After internalization, SCVs can persist within host cells and cause persistent or recurrent infections. Most knowledge on SCVs derives from Staphylococcus species, which are primary models for understanding their phenotypic, genetic, and clinical features (16). Staphylococcus aureus can persist within host cells for extended periods by adopting SCV phenotypes that are reversible upon environmental shifts, thereby contributing to therapy-refractory chronic infections (17). Similarly, in carbapenemase-producing Kp, mucoid to nonmucoid switch occurs via diverse mutations in the capsule locus (K locus/cps), with mutation- and strain-dependent effects on carbapenem susceptibility, biofilm formation, and virulence (15). Phenotypic switching in hvKp is increasingly recognized as an adaptive response. Song et al. reported that carbapenem-resistant hvKp undergoes mucoid-to-nonmucoid switch driven by ISKpn26-mediated inactivation of rmpADC or wcaJ. Hypomucoid variants exhibited reduced virulence in murine sepsis models, decreased survival in normal mouse serum, and enhanced urinary tract persistence with increased adhesion and biofilm formation (18). Consistent with the notion that capsule remodeling shapes host–pathogen interactions, a subset of carbapenemase-producing Kp bloodstream isolates harbored loss-of-function mutations in wcaJ, reducing capsule production while increasing complement binding, opsonophagocytosis, and resistance to complement-mediated killing (19). Recent advances have demonstrated that epigenetic regulation — especially DNA methylation — can drive phenotypic plasticity in bacteria. For instance, DNA adenine methylation (DNA adenine methylase-mediated) has been shown to modulate virulence in Kp genotype K1, affecting serum resistance and in vivo pathogenicity (20). Moreover, a comprehensive methylome analysis of 14 Kp strains revealed multiple adenine (6mA) and cytosine (5mC) methylation motifs, suggesting strain-specific methylation landscapes that may influence replication, transcription, and virulence gene regulation (21). Recent work comparing classical and hypervirulent Kp isolates identified significantly enriched methylation marks (GATC and CCWGG motifs) in virulence loci including capsule synthesis genes in hypervirulent strains, indicating epigenetic signatures correlated with pathogenic phenotype (22). In addition, alterations in metabolic regulatory pathways such as the phosphotransferase system (PTS) have been shown to modulate capsular polysaccharide production and thereby influence phagocytosis resistance in Kp, suggesting that macrophage-induced metabolic pressures could trigger similar transcriptional and post-translational adjustments (23). Collectively, these findings indicate that Kp may undergo coordinated epigenetic, transcriptional, and metabolic reprogramming in response to the macrophage environment, enabling rapid phenotypic switching that modulates capsule production, virulence expression, and phagocytosis resistance.
Another important finding of this study is that phenotypic switching correlates with changes in antibiotic susceptibility. The development of tigecycline resistance in Kpsequence type 23 strains is accompanied by a marked reduction in hypermucoviscosity and virulence. Tigecycline-resistant mutants exhibit decreased CPS production and mucoviscosity, impaired biofilm formation, and reduced survival in human serum, macrophages, and fruit flies, indicating that resistance acquisition comes at the cost of attenuated virulence (24). In hypermucoviscous Kpsequence type 23 strains, colistin resistance leads to reduced hypermucoviscosity, decreased CPS production, and lipid A modifications. These mutants showed impaired survival in human serum and significant fitness costs, indicating a trade-off between resistance and virulence (25). In our study, colistin susceptibility differed among phenotypic variants, with the mixed population exhibiting increased susceptibility, likely reflecting phenotypic heterogeneity. Thus, phenotypic plasticity in hvKp shapes host-pathogen interactions and contributes to antibiotic tolerance and the spread of resistance.
Recent studies have highlighted the clinical significance of capsule regulation in hvKp. For example, Huang et al. demonstrated that serotype-dependent capsule expression governs bloodstream survival and evasion of Kupffer cell clearance (26). Using an ordered transposon mutant library, Mike et al. showed that capsule production and hypermucoviscosity are genetically separable traits with distinct contributions to survival during serum exposure and in vivo fitness (27). Xu et al. showed that RmpA/A2-mediated hypermucoviscosity reduces neutrophil phagocytosis and increases virulence (28).
Taken together, our findings and previous studies indicate that phenotypic switching in hvKp—mucoid and nonmucoid phenotypes, capsule variation, and emergence of SCVs—profoundly influences bacterial morphology, virulence, host–pathogen interactions, and antibiotic susceptibility. These adaptive changes often impose trade-offs, whereby resistance acquisition or enhanced persistence in specific host niches occurs at the cost of attenuated virulence. Recent evidence emphasizes that capsule regulation is central to these processes, positioning phenotypic plasticity as a key driver of hvKp pathogenesis and a major challenge for effective therapeutic intervention.