INTRODUCTION
Carbapenems have long been regarded as reliable therapeutic agents for infections caused by multidrug-resistant Gram-negative bacteria (1, 2). However, the global increase in carbapenem use has been accompanied by a rapid rise in carbapenem-resistant Enterobacterales (CRE), which pose major challenges to clinical management and infection control (2, 3). In Korea, surveillance data indicate a steady increase in CRE notifications since the 2010s, particularly among high-risk populations such as intensive care unit patients, immunocompromised individuals, and those with prolonged hospitalization (4, 5, 6, 7).
Carbapenemase production is the most clinically significant mechanism of carbapenem resistance (8, 9). Carbapenemase- producing Enterobacterales (CPE) disseminate rapidly through plasmid-mediated horizontal transfer, contributing to institutional outbreaks and regional spread (8, 9). The most prevalent carbapenemases worldwide include Klebsiella pneumoniae carbapenemase (KPC), New Delhi metallo-β-lactamase (NDM), and oxacillinase-48-like (OXA-48-like)enzymes, all of which severely limit therapeutic options and are associated with poor clinical outcomes (3, 10, 11).
In Korea, KPC-2-producing Enterobacterales have been consistently reported as the predominant carbapenemase producers, whereas the increasing detection of NDM- and OXA-type carbapenemases reflects expanding genetic diversity (4, 5, 6, 12, 13, 14). Bloodstream CRE infections are of particular concern because they are strongly associated with sepsis, septic shock, and high mortality (15, 16). Despite this clinical importance, long-term regional data describing antimicrobial resistance patterns and carbapenemase distribution among bloodstream CRE isolates remain limited.
Busan is a large metropolitan city with numerous tertiary-care and long-term care hospitals and serves as a major regional referral hub. These characteristics are likely to contribute to the emergence and dissemination of CRE.Therefore, this study analyzed CRE bloodstream isolates collected over a seven-year period to describe antimicrobial resistance patterns, characterize carbapenemase genotypes, and compare regional findings with national epidemiologic trends.
MATERIALS AND METHODS
Bacterial isolates
A total of 609 non-duplicate CRE isolates were recovered from blood cultures at tertiary-care, general, and long-term care hospitals, as well as clinics in Busan, South Korea, between 2018 and 2024.For patients with multiple CRE isolates, only the first isolate per patient during the study period was included in the analysis.Species identification was performed using the Vitek 2 Compact system (bioMérieux, France).
Antimicrobial susceptibility testing
Minimum inhibitory concentrations (MICs) for ertapenem, imipenem, meropenem, and doripenem were determined using Sensititre™ KORN MIC plates (Thermo Scientific, UK) according to the manufacturer’s instructionsand standard laboratory procedures. Antimicrobial susceptibility results were interpreted in accordance with the Compendium of Standard Laboratory Procedures for Infectious Disease Testing issued by the Korea Disease Control and Prevention Agency (KDCA) (17), and susceptibility breakpoints were applied according to the Clinical and Laboratory Standards Institute (CLSI) M100 performance standards for antimicrobial susceptibility testing (18). Quality control was performed using Escherichia coli NCCP 14762 and Pseudomonas aeruginosa NCCP 14570.
Detection of carbapenemase genes
Ten carbapenemase genes (KPC, NDM, OXA, IMP, VIM, GES, SPM, SIM, GIM, and SME) were detected by PCR using the SB-Plex™ CRE Detection Kit (SNB, Korea). Representative amplicons were confirmed by sequencing (Genotech, Korea). Primer sequences are listed in Table 1. Non-CPE isolates were defined as CRE isolates that met phenotypic carbapenem resistance criteria but tested negative for the targeted carbapenemase genes included in the PCR panel.
Table 1.
Primers used for polymerase chain reaction (PCR) detection of carbapenemase genes
| Target gene | Primer name | Sequence (5’→ 3’) | Product size (bp) |
|---|---|---|---|
| IMP |
Forward Reverse |
TGA GCA ATG TAT CTG TAT TC TTA GTT GCT TGG TTT TGA TG | 740 |
| OXA-48 |
Forward Reverse |
TTG GTG GCA TCG ATT ATC GG GAG CAC TTC TTT TGT GAT GGC | 743 |
| VIM |
Forward Reverse |
TGG TCT ACA TGA CCG CGT CT CGA CTG AGC GAT TTG TGT G | 766 |
| NDM |
Forward Reverse |
CAA TAT TAT GCA CCC GGT CG ATC ATG CTG GCC TTG GGG AA | 720 |
| KPC |
Forward Reverse |
ATG TCA CTG TAT CGC GGT CT TTT TCA GAG CCT TAC TGC CC | 893 |
| GES |
Forward Reverse |
GCG CTT CAT TCA CGC ACT AT GCG TAA TCT CTC TCC TGG GC | 753 |
| SME |
Forward Reverse |
AAC GGC TTC ATT TTT G GCT TCC GCA ATA GTT TTA TCA | 830 |
| GIM |
Forward Reverse |
TCG ACA CAC CTT GGT CTG AA AAC TTC CAA CTT TGC CAT GC | 477 |
| SIM |
Forward Reverse |
TAC AAG GGA TTC GGC ATC G TAA TGG CCT GTT CCC ATG TG | 570 |
| SPM |
Forward Reverse |
AAA ATC TGG GTA CTC AAA CG ACA TTA TCC GCT GGA ACA GG | 271 |
| 16S rRNA |
Forward Reverse |
AGA GTT TGA TCM* TGG CTC AG TAC GGY** TAC CTT GTT ACG ACT T | 1465 |
Statistical analysis
Antimicrobial susceptibility and carbapenemase gene detection data were organized using Microsoft Excel 2021 (Microsoft, USA). Differences by year, bacterial species, and type of healthcare institution were analyzed using the chi-square (χ²) test. Temporal trends in major carbapenemase genotypes were assessed using the Cochran-Armitage trend test. A p value < 0.05 was considered statistically significant. All analyses were conducted at the isolate level using only the first isolate per patient. No isolates harboring more than one carbapenemase gene were identified.
RESULTS AND DISCUSSION
Distribution of isolates by healthcare institution type
From 2018 to 2024, a total of 609 CRE bloodstream isolates were collected from 51 healthcare institutions in Busan (Table 2). Most isolates were obtained from tertiary-care hospitals (37.7-57.6%) and general hospitals (33.9-54.7%), together accounting for more than 85% each year. CRE isolation from long-term care hospitals was consistently low (1.7-8.3%), while small numbers were also recovered from community clinics, with a slight increase observed in 2023 (8.8%).
Table 2.
Distribution of Carbapenem-resistant Enterobacterales (CRE) isolates by type of healthcare institution, 2018-2024 (n, %)
These findings indicate that CRE bacteremia predominantly occurs in large healthcare facilities, but smaller institutions also play a role in potential community dissemination. This distribution likely reflects differences in patient populations, disease severity, and diagnostic accessibility across healthcare tiers. Tertiary-care and general hospitals treat critically ill or immunocompromised patients and perform more frequent blood cultures, while long-term care and primary hospitals often refer severe cases.
Similar patterns have been reported in other Korean regional studies, where NDM and OXA-type enzymes were occasionally detected in smaller institutions (4, 5, 6, 12, 13). Furthermore, intensive care units have been identified as key sites for CRE acquisition due to heavy antimicrobial pressure and frequent invasive procedures (7), supporting the concentration of CRE isolates in high-level hospitals observed in this study.
Species distribution of CRE bloodstream isolates
Species analysis showed that K. pneumoniae was the predominant organism, accounting for 81.8% (n=498) of all isolates, followed by E. coli (11.7%, n=71) and Enterobacter spp. (4.6%, n=28) (Table 3). Minor species included Citrobacter spp., Serratia spp., Proteus spp., and Raoultella spp. Year-to-year variation was not statistically significant (p = 0.224).
Table 3.
Distribution of Carbapenem-resistant Enterobacterales (CRE) bloodstream isolates by species in Busan, 2018-2024
This consistent predominance of Klebsiella pneumoniae indicates its major role in CRE bloodstream infections in Busan throughout the study period. The trend aligns with previous Korean multicenter studies (4, 5, 14) and international surveillance reports (2, 3, 11), which have also identified K. pneumoniae as the main pathogen responsible for CRE bacteremia. Given that K. pneumoniae infections are strongly associated with sepsis, septic shock, and high mortality (15, 16), the sustained dominance of this species has important clinical implications for treatment and infection control.
Meanwhile, E. coli and Enterobacter spp. isolates were less frequent but have been reported to harbor NDM- and OXA-type carbapenemases more frequently than K. pneumoniae in regional surveillance and genomic studies, indicating greater genetic diversity and highlighting the need for continued molecular surveillance (19).
Carbapenem resistance profiles
The minimum inhibitory concentration (MIC) distributions for four carbapenems are shown in Table 4 and Fig. 1. Resistance to ertapenem (MIC ≥ 2 µg/mL) was observed in 99.3% of isolates, with 69.3% distributed within the 4-16 µg/mL range. Resistance to imipenem and meropenem (MIC ≥ 4 µg/mL) was identified in 82.0% and 74.8% of isolates, respectively. Resistance to doripenem (MIC ≥ 4 µg/mL) was detected in 58.0% of isolates, while an additional 28.2% of isolates showed MICs of 2 µg/mL, immediately below the resistance breakpoint.
Table 4.
Distribution of minimum inhibitory concentrations (MICs) of four carbapenem agents among Carbapenem- resistant Enterobacterales (CRE) bloodstream isolates
※Resistance (R) breakpoints according to the Clinical and Laboratory Standards Institute (CLSI) M100 performance standards (18); ertapenem ≥2 µg/mL, imipenem, meropenem, and doripenem ≥4 µg/mL.Percentages may not sum to 100% due to rounding to one decimal place.
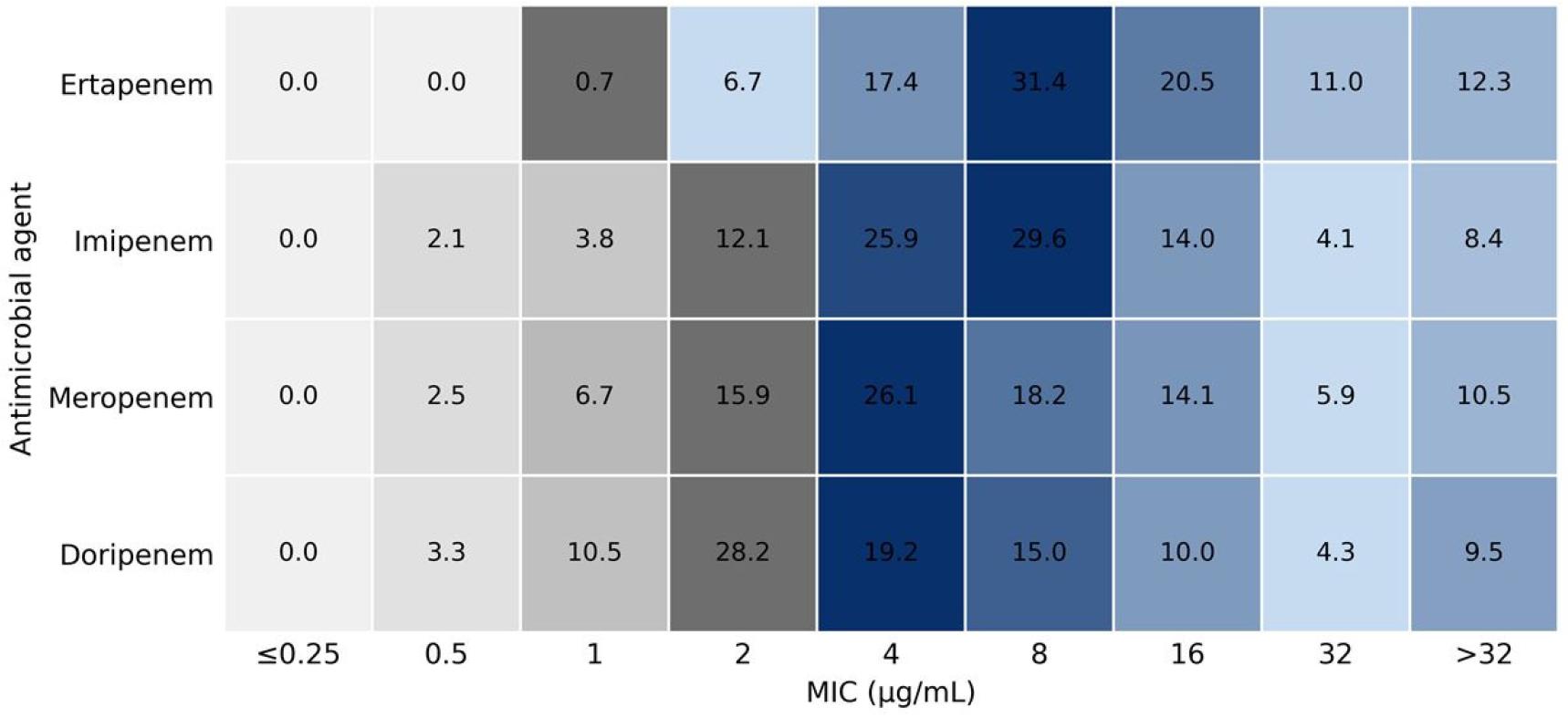
Fig. 1
Distribution of minimum inhibitory concentrations (MICs) of four carbapenem agents among Carbapenem-resistant Enterobacterales (CRE) bloodstream isolates collected in Busan, 2018-2024. Gray indicates the susceptible range (≤1 µg/mL for ertapenem; ≤2 µg/mL for imipenem, meropenem, and doripenem), and blue indicates the resistant range (≥2 µg/mL for ertapenem; ≥4 µg/mL for the other agents). Color intensity corresponds to the proportion of isolates at each MIC value, with darker shades indicating higher frequencies. Breakpoints were interpreted according to Clinical and Laboratory Standards Institute (CLSI) guidelines (18).
Across all four carbapenems, a considerable proportion of isolates demonstrated elevated MIC values (≥ 8 µg/mL), particularly for doripenem and meropenem, indicating that high-level resistance was not restricted to a single carbapenem. This distribution pattern is consistent with previous national surveillance data and international reports showing that decreased susceptibility to ertapenem is often observed prior to the extension of resistance to other carbapenems among CRE isolates (4, 5, 10, 11).
Distribution of carbapenem resistance determinants
Among the 609 isolates, 598 (98.2%) were identified as carbapenemase-producing Enterobacterales (CPE). The remaining isolates, classified as non-CPE (non-carbapenemase-producing Enterobacterales), fulfilled phenotypic carbapenem resistance criteria but were negative for the targeted carbapenemase genes included in the PCR panel. These non-CPE isolates were retained in the analysis as phenotypically carbapenem-resistant isolates lacking PCR-detectable carbapenemase genes. Accordingly, the “non-CPE” category in this study refers to CRE isolates without detectable carbapenemase genes by the assay used, rather than indicating the absence of resistance determinants. These isolates may possess alternative resistance mechanisms, such as porin alterations combined with ESBL or AmpC overexpression, or other mechanisms not covered by the assay. KPC-type enzymes predominated, accounting for 88.2% of all CRE isolates (n = 537), with KPC-2 accounting for 84.9% (n=517) and remaining consistently prevalent throughout the study period (Table 5, Fig. 2). KPC-3 (3.0%) and KPC-4 (0.3%) were detected sporadically. NDM-type enzymes were found in 8.5% of isolates (NDM-1, 6.8%; NDM-5, 1.5%; NDM-6, 0.3%), and OXA-type carbapenemases accounted for 1.5%.
Table 5.
Annual distribution of carbapenem resistance determinants among Carbapenem-resistant Enterobacterales (CRE) bloodstream isolates in Busan, 2018-2024 (n= 609)
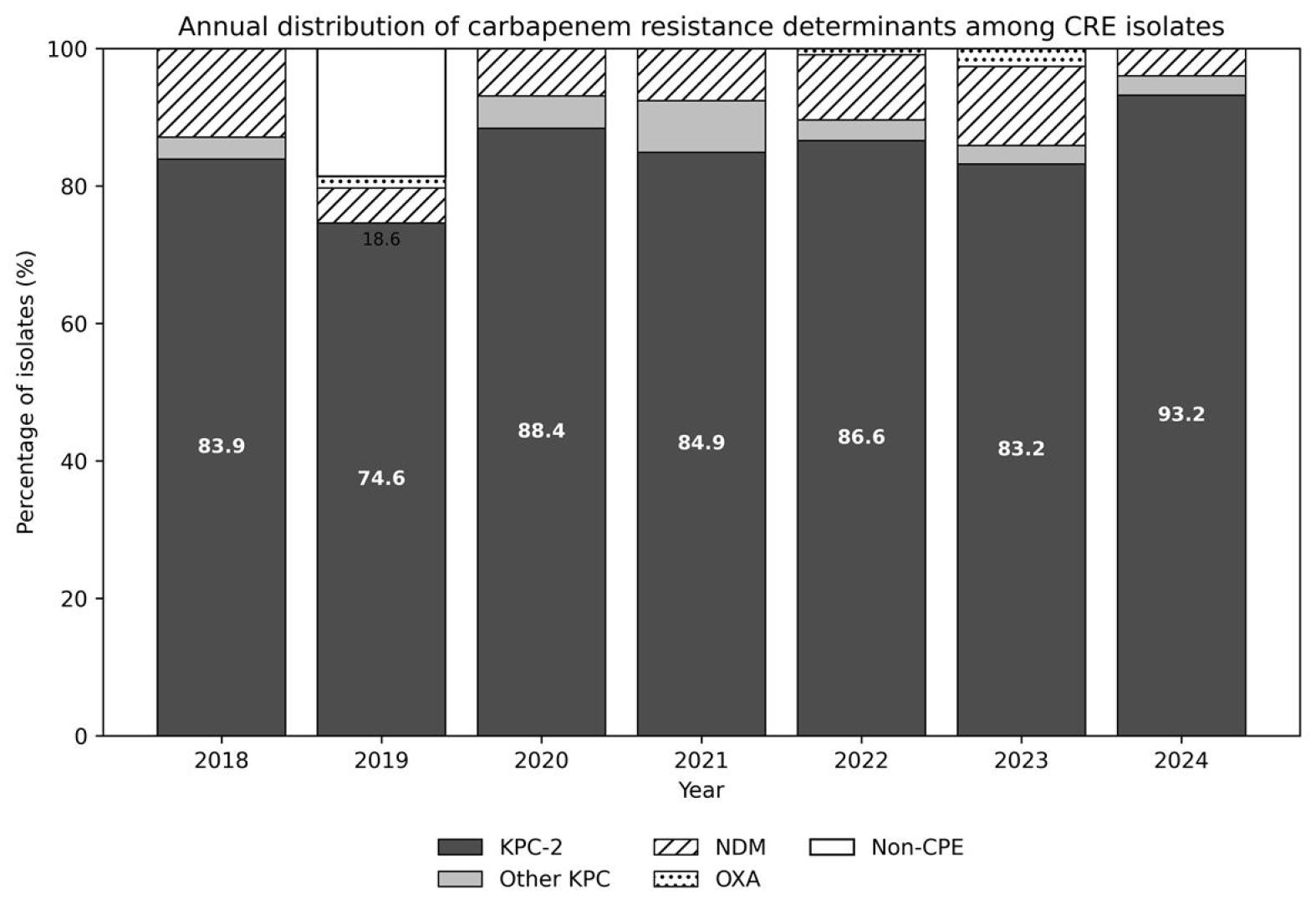
Fig. 2
Annual distribution of carbapenem resistance determinants among Carbapenem-resistant Enterobacterales (CRE) bloodstream isolates in Busan, 2018-2024. Bars represent the percentage distribution of resistance determinants within each year (100% stacked). Percentages were calculated using the total number of CRE isolates identified in each year as the denominator (year-specific n values are presented in Table 2). Each isolate was counted once. Values inside the bars indicate the percentage of KPC-2 isolates. Other-KPC includes KPC-3 and KPC-4. NDM includes NDM-1, NDM-5, and NDM-6. OXA includes OXA-48, OXA-181, and OXA-484. The non-CPE category includes phenotypic CRE isolates negative for the carbapenemase genes targeted in the PCR assay. Percentages may not sum to 100% because of rounding.
Statistical analysis confirmed significant annual differences in genotype distribution (p < 0.001). The consistent dominance of KPC-2 reflects the establishment of a stable regional dissemination pattern within the healthcare network, while sporadic detection of NDM and OXA variants suggests localized outbreaks or plasmid transfer events.
These findings correspond to national surveillance trends showing KPC-2 predominance and emerging NDM variants (4, 5, 19, 20). Globally, geographic heterogeneity in carbapenemase epidemiology has been reported: KPC-type carbapenemases are frequently described in North America and Europe, whereas NDM- and OXA-48-like enzymes are more commonly reported in South Asia and the Middle East, with variable prevalence across East Asia (3, 11, 12).
Species- and healthcare institution-type distribution of carbapenemasegenotypes
Species-specific analysis revealed that K. pneumoniae accounted for 81.8% of isolates and predominantly carried KPC-2 (90.2%). E. coli and Enterobacter spp. showed greater heterogeneity, harboring KPC-3, NDM-1, NDM-5, and OXA-type variants (Fig. 3).
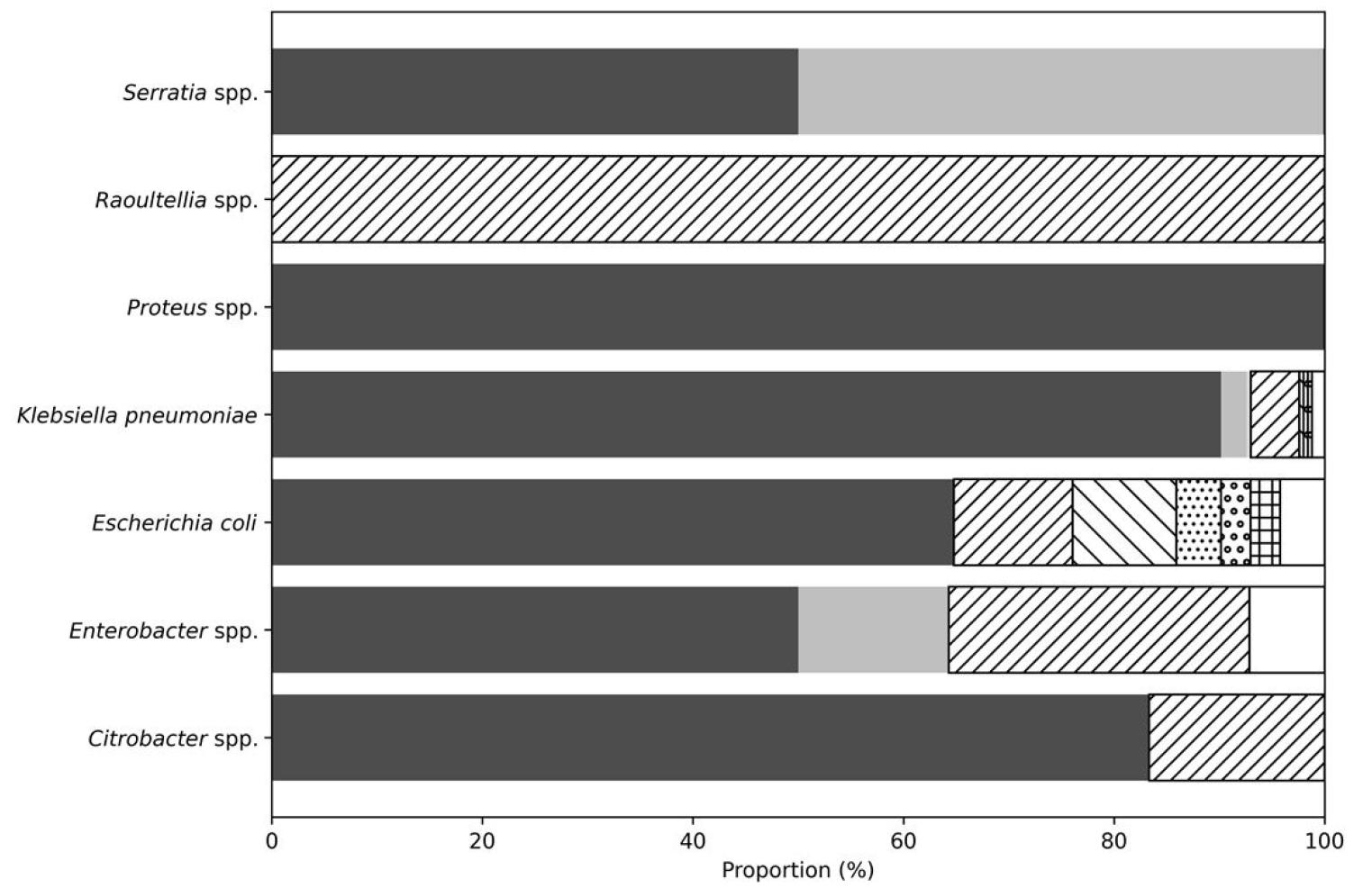
Fig. 3
Distribution of carbapenem resistance determinants by bacterial species among bloodstream isolates in Busan, 2018-2024. Each bar represents the percentage distribution of genotypes within each species (100% stacked). Different fill patterns indicate carbapenemase genotype groups: KPC-2; other KPC (KPC-3 and KPC-4); NDM (NDM-1, NDM-5, and NDM-6); OXA (OXA-48, OXA-181, and OXA-484); and non-CPE.
The differences in genotype distribution by species were statistically significant (χ²-test, p < 0.001). Across healthcare institution types,the annual distribution of carbapenemase genotypes differed (Fig. 4). KPC-2 was most frequent in tertiary-care hospitals (86.3%) and general hospitals (83.3%), whereas long-term care hospitals exhibited a relatively higher proportion of KPC-3 (12.0%), and clinics occasionally yielded NDM-1 (12.0%) and OXA-type isolates (Table 6, Fig. 4).
Table 6.
Distribution of carbapenemase-producing Enterobacterales (CPE) genotypes among bloodstream isolates by healthcare institution type in Busan (2018-2024)
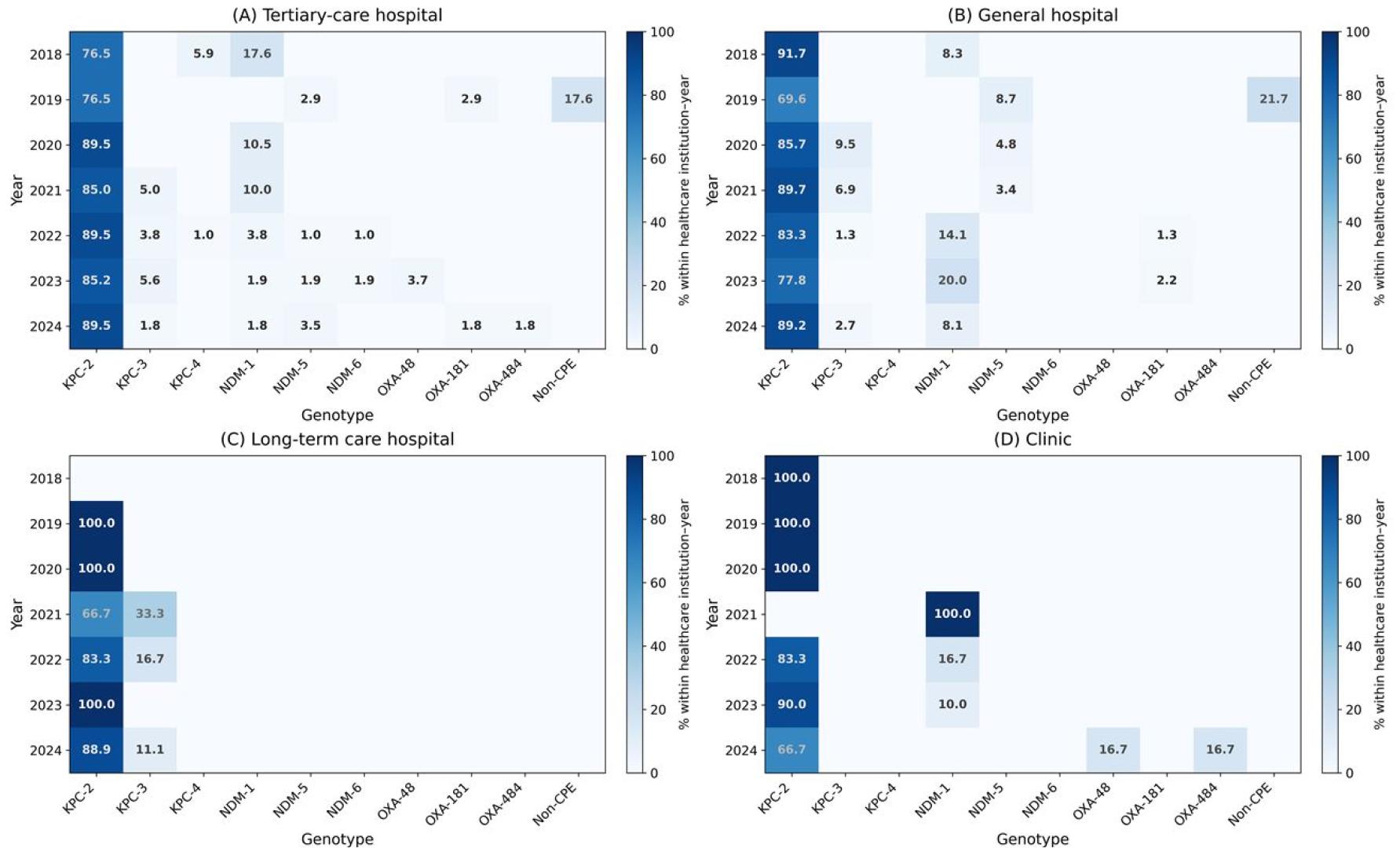
Fig. 4
Annual distribution of carbapenemase genotypes among carbapenem-resistant Enterobacterales (CRE) bloodstream isolates according to healthcare institution type in Busan, 2018–2024. Heatmaps display the percentage of each carbapenemase genotype within each healthcare institution–year stratum. Panels represent (A) tertiary-care hospitals, (B) general hospitals, (C) long-term care hospitals, and (D) clinics. Color intensity increases with higher percentages; only non-zero values are shown.
These findings suggest that tertiary-care hospitals may function as major reservoirs for KPC-2-producing strains, whereas smaller healthcare facilities may contribute to the observed diversity of carbapenemase genotypes through the detection of less common carbapenemase variants. This pattern aligns with previous national and international studies reporting long-term care and rehabilitation facilities as potential sources of emerging carbapenemase variants (1, 2, 4, 7, 20).
Clinical and Epidemiologic Interpretation
This long-term regional surveillance shows the sustained predominance of KPC-2-producing K. pneumoniae as the principal cause of CRE bloodstream infections in Busan, consistent with national surveillance indicating KPC-type carbapenemases as the dominant resistance mechanism in Korea (4, 5, 6, 13, 20). The sporadic detection of NDM- and OXA-type carbapenemases suggests ongoing diversification of resistance genotypes and the potential for dissemination within healthcare networks, including inter-facility patient transfer and healthcare-associated transmission dynamics (2, 7, 8, 21).
A small proportion of isolates (1.8%) were classified as non-CPE, defined as phenotypic CRE isolates lacking PCR-detectable carbapenemase genes. As the PCR assay targeted ten carbapenemase genes, these isolates may reflect alternative resistance mechanisms or genes beyond the assay scope. Similar non-carbapenemase-producing CRE have been increasingly recognized in clinical and molecular studies (21, 22, 23, 24, 25).
This study has limitations. As a single-region surveillance, the findings may not represent nationwide epidemiology. Clinical outcome data were unavailable, and molecular analysis was limited to PCR detection without whole-genome sequencing; therefore, clonal relatedness and novel resistance determinants could not be assessed. These limitations should be considered when interpreting the epidemiological and molecular findings.
CONCLUSION
Our study shows that KPC-2-producing K. pneumoniae was the main cause of CRE bloodstream infections in Busan. Rather than remaining localized, CRE appears to be continuously spreading across healthcare institutions. These findings indicate that sustained molecular surveillance, together with coordinated infection control efforts, should be maintained to limit further dissemination within the regional healthcare system.




