INTRODUCTION
After the discovery of penicillin, the first antibiotic by Alexander Fleming, antibiotics brought about a revolutionary change in modern medicine. Penicillin demonstrated the unprecedented ability to combat bacterial infections effectively, drastically reducing mortality rates from diseases like pneumonia and syphilis, and transforming medical practices by enabling safer surgeries and treatments for previously fatal conditions. This discovery has saved countless lives, changed the paradigm of treating infectious diseases, and contributed to making human life richer and healthier (1). Antibiotics have demonstrated effective therapeutic effects against fatal bacterial infections and have significantly reduced the mortality rate of various bacterial diseases such as endocarditis and bacterial meningitis. They have also shown great efficacy in preventing surgical infections through prophylactic use before and after operations, and have played an important role in protecting immunocompromised and transplant patients. However, there are several drawbacks. Excessive use of antibiotics can lead to toxicities, such as nephrotoxicity associated with an aminoglycoside or colistin. Additionally, antibiotics can affect the commensal flora, leading to opportunistic infections such as Clostridium difficile. The most critical drawback is that antibiotics can induce the development of resistance in bacteria (2). Eventually, the misuse and overuse of antibiotics have led to the emergence of antibiotic resistance, posing a significant threat to human life. This issue has increased morbidity and mortality rates and has become one of the most critical challenges to address in public health (3, 4). In addition, the development of new drugs faces significant challenges due to various limitations, such as economic obstacles. Even when new agents are developed, the emergence of resistance to them is inevitable, making it extremely difficult to combat antibiotic resistance effectively (4). So, the World Health Organization (WHO) has emphasized the urgent need for novel strategies to combat antimicrobial resistance, including the promotion of prudent antibiotic use, the development of new antimicrobial agents, and the exploration of alternative treatments like phage therapy (5).
As a new strategy to combat antimicrobial resistance, the use of bacteriophages —viruses that specifically infect bacteria— in phage therapy is being actively researched. Bacteriophages offer unique advantages such as high specificity for target bacteria, minimizing harm to beneficial microbiota, and the ability to evolve alongside bacteria, reducing the likelihood of resistance development. Although there are some disadvantages, such as the need to select appropriate phages due to their high specificity to their hosts and the potential development of phage resistance, side effects during or after phage application have rarely been reported (6). Phage therapy was first studied in 1919 (7). However, with the discovery of antibiotics, research on phages as a therapeutic approach was largely abandoned. Recently, due to the rise of antibiotic resistance, interest in phage therapy has been reignited, and it is now being actively studied once again. In therapies utilizing phages, when bacteriophages are used in combination with antibiotics, a phenomenon called phage-antibiotic synergy (PAS) may occur (8).
PAS, first studied in 2007, is a phenomenon in which the production of phages increases in the presence of certain antibiotics at sublethal concentrations (9). PAS offers several distinct benefits in addressing bacterial infections, especially antibiotic-resistant strains. First, PAS enables phages and antibiotics to work complementarily, effectively suppressing bacterial growth and replication. For example, streptomycin inhibits bacterial protein synthesis by binding to the 30S ribosomal subunit, specifically the 16S rRNA and a Podovirus such as LUZ7 requires LPS components for infection. (10) In this way, phages and antibiotics can target different bacterial receptors, helping to control bacterial proliferation and antibiotic resistance (11). Moreover, certain antibiotics delay bacterial lysis, inducing morphological changes such as the elongation of bacterial cells. This elongation promotes increased phage replication (12). PAS is also effective in eliminating biofilms, which make treatment significantly more challenging. The matrix of biofilm is degraded by phage enzyme, which increases the permeability of antibiotics (13). PAS also offers significant advantages in controlling bacterial resistance. When phages are used in combination with antibiotics, a broader range of bacteria can be covered. For example, while antibiotics like ciprofloxacin target DNA replication, phages can simultaneously attack bacterial cell walls or membranes, effectively addressing bacterial populations that might otherwise resist single-agent treatments. And by targeting different pathways in bacterial treatment, there is a lower likelihood that bacteria will develop resistance in a short period. Particularly, as bacteria evolve resistance to one agent, changes in their components or functions may make them more vulnerable to other agent (8, 11). In addition, PAS requires lower doses of antibiotics to achieve similar or even better effects, which helps reduce side effects and the development of resistance (11). Furthermore, PAS can reduce bacterial virulence factors by disrupting the mechanisms bacteria use to cause disease, such as toxin production or adhesion to host cells. This reduction not only weakens the bacteria’s ability to cause infections but also enhances the effectiveness of the immune system in clearing the infection (14).
However, despite the many advantages of PAS, much more research is still needed before it can be widely utilized as a treatment. Studies are required to identify optimal phage-antibiotic combinations, assess their efficacy in diverse bacterial strains, and understand the molecular mechanisms underlying their synergy. PAS demonstrates synergy when a specific phage for a specific bacterium and a specific antibiotic are combined. This specificity ensures targeted action, minimizing collateral damage to beneficial microbiota. However, it also poses challenges for clinical application, such as the need for extensive research to identify compatible combinations for diverse bacterial infections. Research on the combination of specific phages and antibiotics that demonstrate synergy is currently ongoing. But, for this research to become practically applicable, the findings need to be consolidated and organized into a cohesive framework that can be effectively implemented in clinical treatments. So, in this review, we will organize and summarize studies on PAS, focusing on which combinations of antibiotics and phages have been effective for specific bacteria. This analysis aims to provide insights into optimizing phage-antibiotic strategies, potentially revolutionizing treatments for antibiotic-resistant infections.
ADVANTAGE OF PAS
a. Antibiotic resensitization
PAS offers several advantages, one of which is the potential to restore bacterial susceptibility to antibiotics. While bacteria resistant to antibiotics alone may not be eradicated, their elimination can be observed when antibiotics are used in combination with phages (15). This effect can occur through various mechanisms, such as sub-lethal doses of antibiotics inducing morphological changes like bacterial filamentation, which enhances phage production and subsequently promotes bacterial death (16). Additionally, phages may target drug efflux pumps, leading to re-sensitization of bacteria to antibiotics (15).
b. Trade-off phenomenon
The term “trade-off” refers to a situation where gaining one benefit comes at the cost of losing another. In the context of PAS, trade-offs can also occur in bacteria. When phages are used therapeutically, bacteria may alter their structures, such as polysaccharides, outer membrane porins, efflux pumps, flagella, and pili, to develop resistance to phages. These structural changes can result in reduced virulence, resensitization to antibiotics, and colonization defects, making bacteria more vulnerable to treatment. For example, Enterococcus faecalis is an opportunistic pathogen that frequently causes hospital-acquired infections and has recently shown increasing resistance to last-resort antibiotics such as daptomycin and vancomycin. Mutations in the Enterococcal polysaccharide antigen (epa) gene cluster of this bacterium confer resistance to lytic phages but also result in a trade-off by increasing sensitivity to antibiotics. A mutation in the epaR gene, for instance, prevents phage adsorption and provides phage resistance, but simultaneously increases susceptibility to daptomycin. Similarly, mutations in the epaX gene can block phage adsorption, yet lead to reduced motility and impaired ability to translocate epithelial cell layers or semisolid surfaces, ultimately decreasing infectivity and survival. Mutations in epaS and epaAC also frequently arise during phage exposure and are associated with enhanced antibiotic susceptibility. Notably, E. faecalis cells lacking epaS show diminished intestinal colonization capacity and are particularly more sensitive to vancomycin (17). Thus, the combination of antibiotics and phages can leverage fitness trade-offs to enhance treatment efficacy and overcome resistance mechanisms.
c. Biofilm removal
The combination of phages and antibiotics is also effective in removing biofilms. Traditional antibiotic therapy alone has struggled to eliminate biofilms, making bacterial eradication challenging. However, when phages are used alongside antibiotics, phage enzymes such as endolysins can degrade the biofilm matrix, increasing the permeability of antibiotics (18). In some cases, the T7 bacteriophage synthesizes a quorum quenching (QQ) lactonase, which hydrolyzes acyl homoserine lactones and inhibits quorum-sensing (QS) activity, thereby suppressing biofilm formation (19). This mechanism enhances the effectiveness of biofilm removal and improves bacterial eradication.
d. Minimizing adverse effects
As mentioned earlier, the use of antibiotics alone can lead to dose-dependent side effects such as nephrotoxicity, as well as disruption of the normal microbiota and the emergence of antibiotic resistance. However, when antibiotics are combined with phages, effective therapeutic outcomes can be achieved with lower doses of antibiotics, thereby minimizing side effects. In particular, phage therapy has rarely been associated with side effects, and although the possibility is low, the emergence of phage resistance can be further reduced when used in combination with antibiotics. Therefore, the combination of antibiotics and phages can serve to effectively minimize the side effects associated with either therapy (11) (Fig. 1).
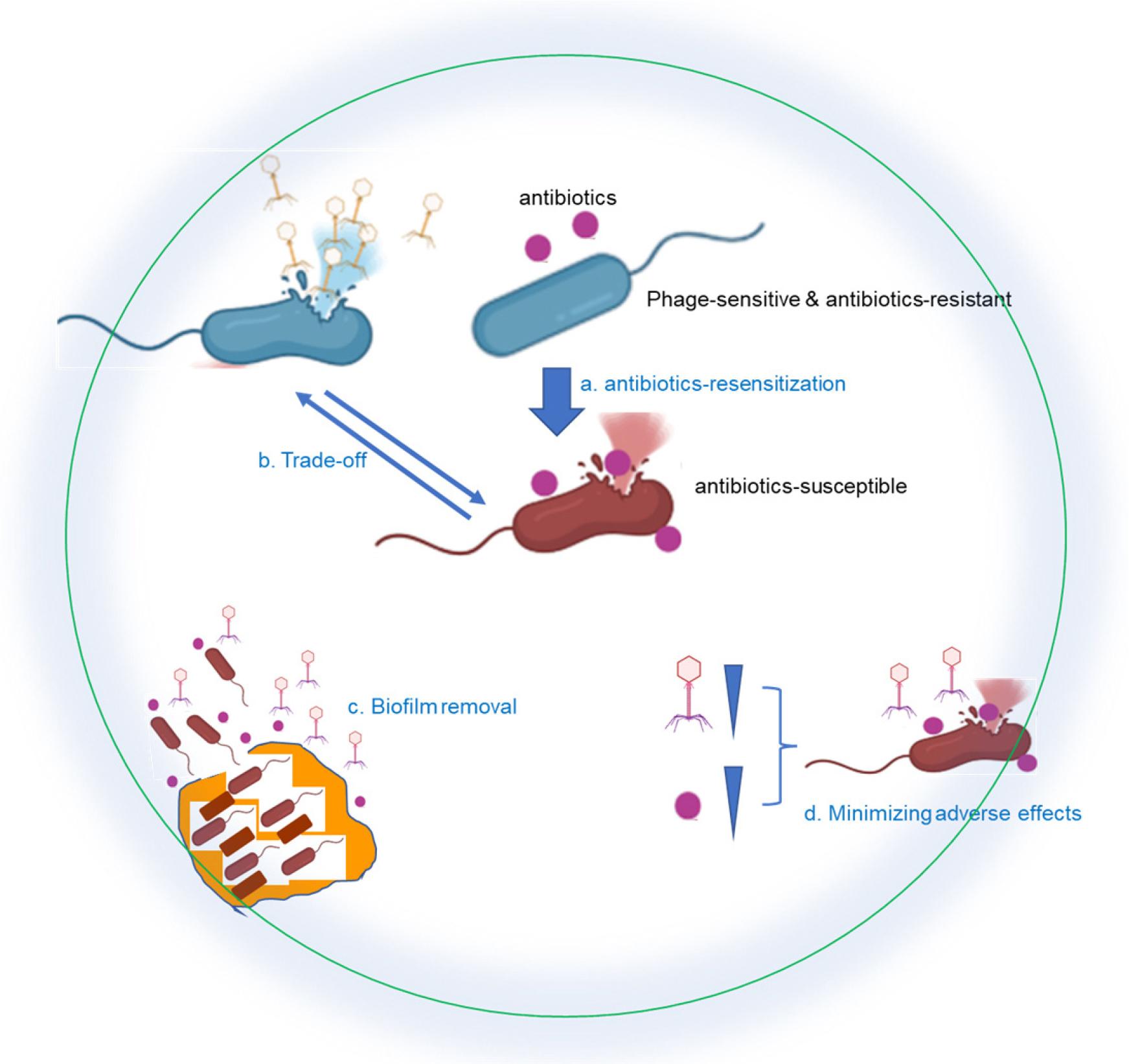
Fig. 1
This diagram illustrates the benefits of phage-antibiotic synergy (PAS) in combating bacterial infections. The top section shows the combination of bacteriophages and antibiotics targeting bacteria. The lower section (labeled a-d) represents specific advantages of PAS: (a) Antibiotic resensitization - The combination of phages and antibiotics restores bacterial susceptibility to antibiotics, enhancing their bactericidal effect. (b) Trade-off effect - Bacteria that develop resistance to phages may become more susceptible to antibiotics or lose virulence factors, reducing their pathogenicity. (c) Biofilm disruption - Phages and antibiotics work together to break down bacterial biofilms, which are otherwise resistant to traditional treatments (illustrated as biofilm structure degradation). (d) Minimizing adverse effects - The combination of phages and antibiotics allows for lower antibiotic doses, reducing toxicity and side effects while maintaining therapeutic efficacy.
PAS REPORTS AGAINST PATHOGENS
Out of the 201 papers initially identified, 83 were selected for analysis, with three of these focusing exclusively on PAS in vivo. The papers included in the review range from the earliest in 2012 to the most recent in 2024. This review covers a total of 13 bacterial species. The most frequently studied bacterium was P. aeruginosa, featured in 20 studies, followed by S. aureus with 19 studies, A. baumannii with 12 studies, and E. coli with 12 studies. Additionally, Klebsiella species were the focus of 8 studies, Enterococcus species appeared in 7 studies, and Salmonella species were included in 3 studies. Furthermore, B. cenocepacia was examined in 2 studies. Lastly, one study each focused on Stenotrophomonas, S. marcescens, Citrobacter, Mycobacterium smegmatis, and Shigella species (Table 1) (20-101). The studies examined a total of 58 overlapping antibiotics, with most bacteria showing both high synergy and high additive/indifferent effects for the same antibiotics.
Table 1.
The number of studies addressing Phage-antibiotic synergy (PAS) for each specific target bacterium
|
PAS (Phage-antibiotic synergy) Study | Target Bacteria (n) | References |
| A. baumannii (12) | 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31 | |
| Klebsiella species (8) K. pneumoniae (6) Klebsiella oxytoca (1) K. quasipneumoniae (1) | 32, 33, 34, 35, 36, 37, 38, 39 | |
| E. coli (12) | 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51 | |
| S. aureus (19) | 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70 | |
| P. aeruginosa (20) | 15, 52, 59, 66, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86 | |
| Enterococcus Species (7) E faecium (6) E. faecalis (2) | 59, 87, 88, 89, 90, 91, 92 | |
| B. cenocepacia (2) | 93, 94 | |
| Stenotrophomonas (1) | 59 | |
| S. marcescens (1) | 95 | |
| Citrobacter (1) | 96 | |
| Mycobacterium smegmatis (1) | 97 | |
| Shigella species (1) S. dysenteriae (1) S. flexneri (1) | 98 | |
| Salmonella Species (3) S. Typhimurium (3) S. Typhi (1) S. Gallinarum (1) S. Blegdam I (1) S. Blegdam II (1) S. Enteritidis (1) | 99, 100, 101 |
These studies employed various methods to evaluate PAS in vitro. The most frequently used method was the time-kill assay, applied in 34 studies. This method evaluates the bactericidal activity of antibiotics by measuring the reduction in viable bacterial count over a specific time frame. By exposing bacteria to antibiotics at defined concentrations and monitoring their survival at intervals, researchers can determine the rate and extent of bacterial killing (102). Following this, checkerboard assays were utilized in 29 studies. Checkerboard assays are two-dimensional, two-agent broth microdilution techniques used to assess the combined effects of antimicrobial agents on bacteria (102). These methods are integral to systematically evaluating PAS. In 13 studies, PAS was evaluated based on bacterial growth. This involved measuring how much bacteria grew over time or after a fixed duration (e.g., 24 hours). In another 11 studies, PAS was assessed using Fractional Inhibitory Concentration (FIC) or Fractional Inhibitory Concentration Index (FICI). FIC is the ratio of an agent’s MIC when used in combination to its MIC when used alone. For instance, FIC for agent A is calculated as (MIC of agent A in combination) / (MIC of agent A). FICI is the sum of the FIC values for two agents (102). In general, checkerboard assays evaluate FIC or FICI together. In addition, various methods, such as evaluating plaque size, have been used to assess PAS. In some cases, a single study employed multiple methods, such as the checkerboard assay and time-kill assay, to evaluate PAS comprehensively.
As with the methods described above, PAS is considered synergistic when the combination shows statistically significant synergy or when the FIC value is less than 0.5. This suggests that the combination of phage and antibiotic is appropriate and results in a synergistic effect. If statistical significance is not demonstrated, or if the FIC value is between 0.5 and 4, the interaction is considered additive or indifferent, indicating that the combination has little or no effect. When the FIC value is 4 or higher, it is interpreted as an antagonistic effect, meaning that the phage-antibiotic combination worsens the situation and is therefore inappropriate (102).
SYNERGY EFFECTS OF PAS REPORTS AGAINST GRAM-NEGATIVE PATHOGENS
- Acinetobacter baumannii
In 12 studies on A. baumannii, the antibiotic showing the most synergy in PAS (14 cases) was meropenem. This was followed by colistin and ampicillin/sulbactam with 9 observed PAS cases each. Tigecycline showed 6 cases of synergy, while rifampicin showed 5 (Table 2). Notably, the A. baumannii strains and phage types used in these 12 studies were all different.
Table 2.
The number of in vitro PAS study showing synergy, indifferent/additive and antagonistic effects for each specific target bacterium. Synergy: The combined effect of two agents is greater than the sum of their individual effects. Indifferent: the effect is similar to the most effective single agent. Additive: The combined effect is equal to the sum of individual effects. Antagonistic effect: The combined effect is less than expected based on individual effects. The exact judgment may vary depending on the criteria of each author
| Target bacteria (n) | Synergy (n) | Ref. | Indifferent / Additive (n) | Ref. | Antagonistic (n) | Ref. | |||
|---|---|---|---|---|---|---|---|---|---|
| A. baumannii (12) |
Carbapenem Penicillin Cephalosporin Monobactam Polymyxin Tetracycline Rifamycin Chloramphenicol Quinolone |
Meropenem (14) Imipenem (3) Ampicillin/Sulbactam (9) Piperacillin (1) Cefotaxime (3) Ceftazidime (2) Cefalotin sodium/sulbactam (1) Aztreonam (1) Colistin (9) Polymyxin B (2) Tigecycline (6) Minocycline (1) Rifampicin (5) Chloramphenicol (1) Ciprofloxacin (2) Levofloxacin (1) | 20, 21, 23, 24, 25, 26, 27, 28, 29, 30, 31 |
Polymyxin Carbapenem Penicillin Quinolone Fosfomycin Aminoglycoside Tetracycline |
Colistin (9) Meropenem (8) Ampicillin/Sulbactam (6) Norfloxacin (3) Ciprofloxacin (2) Fosfomycin (3) Gentamicin (2) Doxycycline (1) | 20, 21, 23, 28 | Aminoglycoside | Gentamicin (1) | 23 |
| K. pneumoniae (6) |
Mitomycin Carbapenem Cephalosporin Polymyxin Quinolone Aminoglycoside Chloramphenicol |
Mitomycin C (4) Meropenem (3) Imipenem (2) Ceftazidime (1) Colistin (2) Ciprofloxacin (2) Levofloxacin (2) Kanamycin (1) Chloramphenicol (1) | 32, 33, 34, 35, 36 |
Aminoglycoside Tetracycline Polymyxin Rifamycin Quinolone Mitomycin Trimethoprim |
Amikacin (3) Tigecycline (2) Polymyxin B (1) Rifampicin (1) Ciprofloxacin (1) Mitomycin C (1) Trimethoprim (1) | 32, 34, 35, 36 | |||
| K. oxytoca (1) | Aminoglycoside |
Gentamicin (1) Amikacin (1) Tobramycin (1) | 38 | ||||||
| K. quasipneumoniae (1) |
Chloramphenicol Aminoglycoside |
Chloramphenicol (1) Neomycin sulfate (1) | 39 | ||||||
| E. coli (12) |
Cephalosporin Carbapenem Penicillin Vancomycin Quinolone Polymyxin Aminoglycoside Fosfomycin Tetracycline Macrolide Rifamycin Chloramphenicol Trimethoprim Mitomycin |
Ceftazidime (8) Cephalexin (4) Cefotaxime (1) Cefaperazone (1) Meropenem (3) Ampicillin (1) Piperacillin (1) Vancomycin (1) Ciprofloxacin (8) Nalidixic acid (1) Colistin (4) Polymyxin B (1) Amikacin (4) Kanamycin (2) Gentamicin (1) Fosfomycin (3) Tetracycline (2) Tigecycline (1) Azithromycin (1) Rifampicin (1) Chloramphenicol (1) Trimethoprim (1) Mitomycin C (1) | 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51 |
Tetracycline Polymyxin Carbapenem Penicillin Cephalosporin Chloramphenicol Aminoglycoside Quinolone Fosfomycin Trimethoprim |
Tigecycline (7) Tetracycline (2) Colistin (5) Meropenem (5) Ampicillin (1) Cefotaxime (1) Cefixime (1) Ceftazidime (1) Chloramphenicol (4) Kanamycin (4) Streptomycin (3) Amikacin (2) Ciprofloxacin (3) Levofloxacin (1) Oxolinic acid (1) Fosfomycin (2) Trimethoprim (2) | 41, 42, 43, 45, 46, 48, 51 | Quinolone | Ciprofloxacin (1) | 43 |
| S. aureus (18) |
Cyclic lipopeptide Cephalosporin Penicillin Vancomycin Fosfomycin Quinolone Macrolide Lincosamide Tetracycline Polymyxin Chloramphenicol Aminoglycoside Oxazolidinone Rifamycin Combination |
Daptomycin (12) Ceftaroline (7) Cefoxitin (2) Cefotaxime (1) Cefazolin (1) Oxacillin (1) Vancomycin (5) Fosfomycin (2) Ciprofloxacin (5) Erythromycin (5) Azithromycin (3) Clarithromycin (1) Clindamycin (4) Tetracycline (4) Doxycycline (1) Polymyxin B (2) Chloramphenicol (1) Gentamicin (1) Linezolid (1) Rifampin (1) DAP + CPT (12) | 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70 |
Cyclic lipopeptide Cephalosporin Penicillin Vancomycin Fosfomycin Aminoglycoside Quinolone Rifamycin Tetracycline Chloramphenicol Oxazolidinone |
Daptomycin (8) Ceftaroline (5) Cefazolin (2) Amoxicillin (3) Oxacillin (2) Dicloxacillin (1) Vancomycin (4) Fosfomycin (1) Gentamicin (4) Ciprofloxacin (3) Rifampin (2) Tetracycline (2) Chloramphenicol (1) Linezolid (1) | 53, 55, 56, 57, 58, 60, 62, 66, 68, 69 |
Oxazolidinone Tetracycline |
Linezolid (2) Minocycline (2) | 59 |
| P. aeruginosa (20) |
Quinolone Carbapenem Cephalosporin Penicillin Monobactam Fosfomycin Aminoglycoside Polymyxin Chloramphenicol Combination |
Ciprofloxacin (35) Meropenem (25) Ceftazidime (21) Cefepime (7) Cefotaxime (5) Cefoperazone (5) Cefozopran (4) Latamoxef (4) Sulbactam/ Cefeperazone (3) Ceftriaxone (1) Piperacillin (4) Aztreonam (7) Aztreonam lysine (2) Fosfomycin (3) Gentamicin (21) Amikacin (6) Tobramycin (3) Colistin (4) Chloramphenicol (1) COL + CIP (1) COL + MEM (1) | 15, 52, 59, 66, 71, 72, 73, 74, 75, 77, 78, 79, 80, 81, 82, 84, 85, 86 |
Aminoglycoside Polymyxin Quinolone Carbapenem Monobactam Cephalosporin Penicillin Fosfomycin Tetracycline Chloramphenicol Sulfonamide Macrolide |
Tobramycin (15) Gentamicin (13) Amikacin (9) Kanamycin (1) Colistin (14) Ciprofloxacin (12) Levofloxacin (7) Meropenem (12) Imipenem (7) Aztreonam (9) Ceftazidime (7) Cefpodoxime (7) Cefotiam (7) Cefmetazole (7) Sulbactam/ Cefeperazone (4) Cefepime (3) Cefozopran (3) Cefotaxime (3) Latamoxef (3) Cefoperazone (2) Piperacillin (3) Fosfomycin (4) Minocycline (7) Tetracycline (2) Chloramphenicol (7) Sulfamethoxazole/Trimethoprim (7) Erythromycin (1) | 15, 66, 71, 72, 73, 74, 75, 76, 77, 78, 80, 81, 82, 83, 85 |
Macrolide Quinolone |
Azithromycin (2) Ciprofloxacin (1) | 59, 72 |
| E. faecium (6) |
Penicillin Cephalosporin Vancomycin Cyclic lipopeptide Oxazolidinone Combination |
Ampicillin (24) Ceftaroline (2) Vancomycin (1) Daptomycin (5) Linezolid (1) DAP + AMP (13) DAP + CPT (7) DAP + ERT (2) | 59, 87, 88, 89, 90, 91 |
Cyclic lipopeptide Penicillin Cephalosporin Carbapenem Combination |
Daptomycin (31) Ampicillin (11) Ceftaroline (3) Ertapenem (3) DAP +AMP (22) DAP + CPT (1) DAP + ERT (3) | 89, 90, 91 |
Oxazolidinone Tetracycline |
Linezolid (2) Minocycline (2) | 59 |
| E. faecalis (2) |
Vancomycin Oxazolidinone |
Vancomycin (2) Linezolid (1) | 87, 92 | ||||||
| B. cenocepacia (2) |
Quinolone Carbapenem Tetracycline Trimethoprim |
Ciprofloxacin (2) Meropenem (2) Tetracycline (2) Trimethoprim (1) | 93, 94 | ||||||
| Stenotrophomonas (1) | Cephalosporin | Ceftazidime (2) | 87 | ||||||
| S. marcescens (1) |
Penicillin Carbapenem |
Ampicillin/Sulbactam (1) Meropenem (1) | 95 | ||||||
| Citrobacter (1) |
Carbapenem Penicillin Cephalosporin Fosfomycin Polymyxin Aminoglycoside Tetracycline |
Meropenem (1) Carbenicillin (1) Cefotaxime (1) Cefepime-tazobactam (1) Fosfomycin (1) Colistin (1) Gentamicin (1) Tigecycline (1) | 96 | ||||||
| Mycobacterium smegmatis (1) | Rifamycin | Rifampicin (1) | 97 | ||||||
| S. dysenteriae (1) |
Cephalosporin Macrolide Aminoglycoside Novobiocin Polymyxin |
Cefotaxime (1) Cefoxitin (1) Cephalothin (1) Erythromycin (1) Gentamicin (1) Cardelmycin (1) Polymyxin B (1) | 98 | ||||||
| S. flexneri (1) |
Cephalosporin Macrolide Aminoglycoside Novobiocin Polymyxin |
Cefotaxime (1) Cefoxitin (1) Cephalothin (1) Erythromycin (1) Gentamicin (1) Cardelmycin (1) Polymyxin B (1) | 98 |
Chloramphenicol Tetracycline Cephalosporin Macrolide Aminoglycoside Novobiocin Polymyxin |
Chloramphenicol (2) Tetracycline (2) Cefotaxime (1) Cefoxitin (1) Cephalothin (1) Erythromycin (1) Gentamicin (1) Cardelmycin (1) Polymyxin B (1) | 99 | |||
| S. Typhimurium(3) |
Aminoglycoside Quinolone Penicillin |
Kanamycin (2) Ciprofloxacin (1) Ampicillin (1) | 99, 100, 101 | ||||||
| S. Typhi(1) | Aminoglycoside | Kanamycin (1) | 99 | ||||||
| S. Gallinarum (1) | Aminoglycoside | Kanamycin (1) | 99 | ||||||
| S. Blegdam I (1) | Aminoglycoside | Kanamycin (1) | 99 | ||||||
| S. Blegdam II (1) | Aminoglycoside | Kanamycin (1) | 99 | ||||||
| S. Enteritidis (1) | Aminoglycoside | Kanamycin (1) | 99 | ||||||
In our laboratory, we studied the synergy between Phage vB_AbaSi_W9 and antibiotics targeting A. baumannii(21). Five strains were tested: A. baumannii ATCC1978, A. baumannii KBN10P04948 (ST191), A. baumannii LIS2013230 (ST208), A. baumannii KBN10P05982 (ST369), and A. baumannii KBN10P05231 (ST451). The antibiotics tested included meropenem, colistin, ampicillin/sulbactam, tigecycline, and rifampicin. Tigecycline and rifampicin exhibited synergy across all strains, while colistin showed indifferent results in all strains except for ST369. Meropenem showed synergy with ATCC1978 and ST369, while ampicillin/sulbactam demonstrated synergy with ATCC1978, ST369, and ST451 (21).
- Klebsiella species
Klebsiella species were studied in a total of 8 papers, with 6 focusing on K. pneumoniae and 2 on K. oxytoca and K. quasipneumoniae, respectively. Mitomycin C demonstrated the most synergy for K. pneumoniae, with 4 PAS cases observed, followed by meropenem with 3 PAS cases. Additionally, imipenem, colistin, ciprofloxacin, and levofloxacin each showed 2 PAS cases. Several other antibiotics also displayed synergy (Table 2). All studies, except for 2, used different strains and phages. Phage vB_KpnM-VAC13 was studied in two papers. Pacios et al. (2021) investigated PAS using K. pneumoniae K2534 and K3325 with combinations of phage vB_KpnM-VAC13 and mitomycin C, as well as phage vB_KpnM-VAC13 and imipenem. (33) Pacios et al. (2024) studied PAS using K. pneumoniae K2534, K3325, and ATCC10031 in combinations of phage vB_KpnM-VAC13 and mitomycin C. (34) Interestingly, the Pacios’s 2021 study demonstrated synergy in all combinations, with phage and mitomycin C preventing resistance in K2534. However, the Pacios’s 2024 study found that the combination failed to prevent resistance in K2534. This highlights the need for further research to identify optimal phage-antibiotic combinations and better understand their mechanisms of action.
- Escherichia coli
In the 12 studies on E. coli, ceftazidime and ciprofloxacin exhibited the most synergy, with 8 PAS each, followed by cephalexin, colistin, and amikacin with 4 PAS each. Several other antibiotics also showed synergy (Table 2). Two studies examined phage T4. Ryan et al. demonstrated synergistic effects with phage T4 and cefotaxime against E. coli 11303, while Scanlan et al. found no synergy when combining phage T4 with chloramphenicol, ciprofloxacin, streptomycin, or tetracycline against E. coli K12 MG1655. Two additional studies explored phage HK97, both targeting E. coli K-12. Despite using different antibiotics, nalidixic acid, oxolinic acid, levofloxacin, mitomycin C, trimethoprim, and others exhibited synergy. (40, 41) In studies targeting previous bacterial strains, antibiotics showing high synergy often also exhibited high additive/indifferent effects. For E. coli, tigecycline demonstrated only 1 case of synergy but 7 cases of additive/indifferent effects. This suggests that tigecycline may not be suitable for PAS involving E. coli.
In our laboratory, we studied the synergy between Phages EC.W1-9 and EC.W15-4 and antibiotics against ESBL-producing and CREC isolates (103). While some PAS was observed with each phage individually combined with colistin, the overall effect was limited. However, when using a phage combination (EC.W1-9 and EC.W15-4) with antibiotics, greater PAS was observed. Among the 12 isolates of ESBL-producing and CREC tested, the phage combination with colistin, meropenem, and tigecycline demonstrated significant synergy. In contrast, amikacin and ciprofloxacin did not yield improved synergistic effects.
- Pseudomonas aeruginosa
A total of 20 studies focused on P. aeruginosa, with ciprofloxacin showing the most synergy, reported in 35 PAS cases, followed by meropenem (25 cases), ceftazidime, and gentamicin (21 cases each). Various other antibiotics also showed synergy (Table 2). The most studied strain was P. aeruginosa PAO1, examined in 9 studies. Except for two studies using phage EPA1, all studies used different phages. Ciprofloxacin was investigated in 6 PAO1 studies, showing synergy in 5 cases except for its combination with phage KPP22. (71, 75, 77, 83, 85, 86) Meropenem demonstrated consistent synergy across 5 studies, (71, 75, 77, 85, 86) and gentamicin showed synergy in 4 out of 6 studies, particularly with phage EPA1 and phage EPA1+phage SAFA. (52, 71, 77, 83, 84, 86) Two studies focused on P. aeruginosa PA14, examining phage NP1+NP3 and phage JG024 in combination with antibiotics. (66, 73) Both studies demonstrated synergy with ciprofloxacin and indifferent effects with gentamicin. Phage NP1+NP3 showed synergy with ceftazidime, while phage JG024 demonstrated synergy with ceftriaxone. Similarly, P. aeruginosa 10266 and phage EM were investigated in two studies, but no overlapping antibiotics were tested. (59, 79) These studies reported synergy with ciprofloxacin, meropenem, and gentamicin.
In PAS studies targeting Gram-negative pathogens, various antibiotics demonstrated synergistic effects depending on the specific pathogen. Among them, antibiotics from the carbapenem, cephalosporin, and quinolone classes—particularly ciprofloxacin—showed the most frequent synergy. Carbapenems and cephalosporins exert their antibacterial effects by inhibiting bacterial cell wall synthesis. Notably, other antibiotics with the same mechanism, such as penicillins and monobactams, also occasionally exhibited synergy, suggesting that antibiotics targeting cell wall synthesis may be particularly effective in PAS against Gram-negative bacteria. Additionally, antibiotics with other mechanisms, such as ciprofloxacin (which inhibits DNA synthesis) and aminoglycosides (which inhibit protein synthesis), also demonstrated considerable synergy.
SYNERGY EFFECTS OF PAS REPORTS AGAINST GRAM-POSITIVE PATHOGENS
- Staphylococcus aureus
Among the 18 studies on S. aureus, the combination of daptomycin and daptomycin + ceftaroline showed the most synergy, with 12 PAS cases reported. This was followed by ceftaroline (7 cases), vancomycin, ciprofloxacin, and erythromycin (5 cases each). Additional synergy was observed with antibiotics like clindamycin and tetracycline (Table 2). The most frequently studied phage for S. aureus was Phage Sb-1, used in 6 studies. Four studies used Phage Sb-1 alone, while two combined it with other phages (e.g., Intesti13 or Intesti13 + Romulus). Combining Phage Sb-1 with daptomycin resulted in 7 synergistic and 1 indifferent PAS out of 8 cases. However, phage combinations showed limited synergy with daptomycin, achieving only 1 case of synergy. By contrast, the combination of daptomycin and ceftaroline without phages consistently showed synergy in 6 cases. The 4 studies using Phage Sb-1 commonly targeted S. aureus D712. However, studies by Kebriaei et al. (2020, 2022) and Kunz Coyne et al., all focusing on S. aureus D712 with Phage Sb-1, yielded differing results. (55, 56, 59) Kebriaei et al. (2020) observed synergistic effects with vancomycin and daptomycin, while ceftaroline and cefazolin showed indifferent effects. (56) Kebriaei et al. (2022) found synergy only with DAP + CPT but indifferent effects with daptomycin and ceftaroline. (55) Kunz Coyne et al. reported synergy with daptomycin and ceftaroline, but antagonism with linezolid and minocycline. (59) The differences in outcomes, especially for daptomycin and ceftaroline, may stem from Kebriaei et al. (2022) focusing on biofilm models, unlike the Kebriaei’s 2020 study. Similarly, ciprofloxacin showed varying results in two studies targeting S. aureus KACC13236 with Phage SA11. These findings underscore the importance of identifying appropriate combinations tailored to specific treatment scenarios and suggest the need for further research (53, 54).
- Enterococcus faecium
E. faecium was studied in six papers, with ampicillin (AMP) showing the most synergy, reported in 24 PAS cases. This was followed by daptomycin + ampicillin (13 cases), daptomycin + ceftaroline (7 cases), and daptomycin alone (5 cases) (Table 2). The most studied strains were E. faecium R497 (5 studies) and HOU503 (4 studies). R497 is a daptomycin-resistant clinical strain and HOU503 is a vancomycin-resistant, daptomycin-susceptible dose dependent (SDD) clinical strain (90). In studies targeting E. faecium R497, phages NV-497 and phage 113 were investigated in three studies each, while phage NV-503-01 was studied in two. Phage NV-497 alone showed synergy with daptomycin, ceftaroline, DAP+AMP, and DAP+CPT (59, 89, 90). phage 113 exhibited synergy with daptomycin, ampicillin, DAP+AMP, DAP+CPT, and DAP+ertapenem (91, 92). Phage NV-503-01 synergized with DAP+CPT (89). Across combinations involving NV-497, ATCC 113, NV-503-01, and NV-503-02, ampicillin showed the most synergy, followed closely by DAP+AMP (90). For E. faecium HOU503, phage 113 was studied in three papers, NV-503-01 in two, and NV-497 in one. Phage 113 showed indifferent effects with daptomycin, ampicillin, ceftaroline, and ertapenem but demonstrated synergy with DAP+CPT (2 cases), DAP+AMP, and DAP+ertapenem (1 case each). (91, 92) Phage NV-503-01 exhibited synergy with daptomycin and ceftaroline, while phage combinations showed the most synergy with ampicillin (59, 90).
In PAS studies targeting Gram-positive pathogens, various antibiotics showed synergistic effects depending on the specific strain. Among them, daptomycin (a cyclic lipopeptide), ceftaroline (a cephalosporin), and ampicillin (a penicillin) exhibited particularly strong synergy. Daptomycin acts on the bacterial cell membrane, altering its permeability and ultimately leading to cell death, while penicillins and cephalosporins exert their antibacterial effects by inhibiting cell wall synthesis. Against Gram-positive bacteria, antibiotics that target the cell membrane or cell wall have demonstrated significant PAS activity. Notably, combinations of such antibiotics have often resulted in synergistic effects.
In addition, various bacteria showed synergy with the following antibiotics (Table 2).
ANTAGONISTIC EFFECTS OF PAS
For A. baumannii, there was one study by Grygorcewicz et al. that reported an antagonistic effect in a PAS experiment (23). In this study, the combination of phage vB_Aba-4 and gentamicin against A. baumannii PUM 45202 showed this antagonistic interaction. Overall, there were three studies on gentamicin-based PAS for A. baumannii, and aside from the one with antagonism, the results of the other two studies were indifferent. This suggests that combining phage and gentamicin may be less effective in treating A. baumannii, indicating the need for further research what factor(s) influence the antagonistic effect between PUM 45202 phage and gentamicin.
In a study on E. coli, Gu Liu et al. demonstrated an antagonistic effect in a PAS experiment (43). This result was observed when combining phage HP3 with ciprofloxacin against ExPEC JJ2528. Ciprofloxacin is one of the antibiotics that demonstrates the most synergy in PAS studies targeting E. coli, making its antagonistic effect particularly significant. Ciprofloxacin, a quinolone antibiotic, inhibits DNA gyrase, thereby blocking DNA replication (43). When used in combination with phages, the antibiotic’s inhibition of bacterial DNA replication can also interfere with the phage’s DNA replication process, potentially leading to antagonism. However, in the study by Gu Liu et al., synergy was observed at certain doses of ciprofloxacin or phage concentrations, showing that both synergy and antagonism can occur depending on the conditions. This suggests that ciprofloxacin can still be used in combination with phages for the treatment of E. coli, but careful evaluation of the phage-antibiotic combinations and their concentrations is crucial before treatment to avoid antagonistic effects.
In studies targeting S. aureus, antagonistic effects were observed with Linezolid and Minocycline, each showing two instances of antagonism (59). These results were reported by Kunz Coyne et al., with S. aureus D712 and S. aureus 684, respectively, where Phage Sb-1 was combined with Linezolid and Minocycline. Linezolid, an oxazolidinone antibiotic, works by inhibiting protein synthesis through binding to the 50S ribosomal subunit, while Minocycline, a tetracycline-class antibiotic, acts on the 30S ribosomal subunit to similarly block protein synthesis. These antibiotics can interfere not only with bacterial protein synthesis but also with phage replication and production, leading to the observed antagonistic effects when used together with phages.
In studies targeting E. faecium, similar to S. aureus, antagonistic effects were observed with Linezolid and Minocycline (59). These findings were also reported by Kunz Coyne et al. Specifically, this antagonism occurred when E. faecium R497 was combined with phage NV-497 and when E. faecium HOU503 was combined with phage NV-503-01.
P. aeruginosa exhibited antagonistic effects with two antibiotics, reported in a total of three instances. Two cases were observed with azithromycin, and one case was reported with ciprofloxacin. Chang et al. demonstrated antagonism when combining phage PEV20 with ciprofloxacin against P. aeruginosa LESB58 (72). Phage PEV20 is completely resistant to P. aeruginosa LESB58, which could explain the results. Ciprofloxacin was the antibiotic with the most observed synergy against P. aeruginosa in this review. Therefore, the occurrence of antagonistic effects with ciprofloxacin presents a critical issue that must be thoroughly addressed for future PAS applications. There are two additional studies involving phage PEV20, and unlike the Chang’s research, these studies demonstrated a synergistic effect of the combination of phage PEV20 and ciprofloxacin against P. aeruginosa FADD1-PA001 and P. aeruginosa JIP865 (80, 81). To avoid such antagonistic effects, it is crucial to select the appropriate phage, and further research is needed to address this issue. Kunz Coyne et al. also reported antagonism when combining phage 14207 with azithromycin for P. aeruginosa 9010 and phage EM with azithromycin for P. aeruginosa 10266 (59).
PAS REPORTS IN VIVO
Several studies have evaluated PAS in vivo to determine its effectiveness in improving survival rates using various models. A total of 13 studies were conducted: 5 used mice, 4 used Galleria mellonella, and 2 each used rats and zebrafish. The most frequently evaluated bacterium was A. baumannii (5 studies), followed by K. pneumoniae (4 studies), S. aureus (3 studies), and Klebsiella oxytoca and P. aeruginosa (1 study each) (Table 3).
Table 3.
In vivo studies on phage-antibiotic synergy (PAS) against bacterial infections. Different animal models (Galleria mellonella, mice, rats, zebrafish) were used to evaluate the efficacy of phage and antibiotic combinations
| Animal model | Target bacteria | Phage | Antibiotics | Effect | Ref. |
|---|---|---|---|---|---|
| Galleria mellonella | A. baumannii Ab177_GEIH-2000 | Phage Ab105-2phi∆CI | Imipenem | Yes | 20 |
| Mice | A. baumannii KBN10P04948 (ST191) | phage vB_AbaSi_W9 | Rifampicin | Yes | 21 |
| Mice | A. baumannii AB900 | phage øFG02 | Ceftazidime | Yes | 22 |
| Zebrafish | A. baumannii 2023 | Phage pB23 | Meropenem | Yes | 24 |
| Zebrafish | A. baumannii 4015 | Phage YC#06 | Chloramphenicol + Imipenem + Cefotaxime | Yes | 25 |
| Mice | K. pneumoniae KBN10P07398 | Phage vB_KpnM_W17 | Ciprofloxacin | Yes | 32 |
| Galleria mellonella | K. pneumoniae K2534 | Phage vB_KpnM-VAC13 | Mitomycin C | Yes | 33 |
| Imipenem | Yes | ||||
| K. pneumoniae K3325 | Mitomycin C | Yes | |||
| Imipenem | Yes | ||||
| Mice | K. pneumoniae 2058 | Phage cocktail P7 (Phage p52, p60, p67, p79, p85, p28, p61) | Colistin | Yes | 37 |
| Galleria mellonella | Klebsiella oxytoca FK-8388 | Phage vB8388 | Gentamicin | Yes | 38 |
| Rat | S. aureus ATCC51650 | Phage Sa87 | Clindamycin |
Yes Yes | 62 |
| Azithromycin | |||||
| Galleria mellonella | S. aureus (MRSA) SA-28 | Phage vB_Sau_S90 + Phage vB_Sau_S165 | Oxacillin | Yes | 64 |
| S. aureus (MRSA) SA-90 | |||||
| S. aureus (MRSA) SA-165 | |||||
| Rat | S. aureus (MRSA) AW7 | Phage K + Phage 3A | Daptomycin | No | 67 |
| Mice | P. aeruginosa PAO1 | Phage PAM2H | Ceftazidime | Yes | 15 |
The combinations of bacteria, phages, and antibiotics are detailed in Table 3. Among these, the only case without a significant survival rate improvement was in the rat model with MRSA AW7 treated with phage K + phage 3A and daptomycin (52). However, the other 12 studies confirmed the efficacy of PAS. These in vivo experiments validate PAS’s therapeutic potential and its applicability in real treatments. Although further studies are required for human applications, this review highlights the promising prospects of PAS in clinical settings.
LESSONS FROM PAST RESEARCHES
As previously discussed, even when using the same antibiotic against the same bacterial strain, the type of phage can result in different outcomes, including synergy, indifference, or even antagonistic effects. Therefore, finding the appropriate combination of phages and antibiotics and evaluating PAS before treatment is essential for its effective therapeutic application.
In this study, antibiotics that exhibited antagonistic effects were commonly found to have mechanisms that inhibit DNA or protein synthesis. Since phages rely on the host bacteria’s enzymes to replicate their DNA and produce proteins, these antibiotics may have interfered with the replication process of the phages. Thus, further research is needed to explore the mechanisms and optimize the use of these antibiotics in PAS treatments. Precise evaluation before clinical application is also crucial.
This research is expected to provide significant insights into selecting appropriate antibiotics for PAS in future studies and therapeutic applications.
With recent advancements in bioinformatics, the development of programs predicting interactions between bacteria and phages has significantly improved. These advancements have made predicting phage-host interactions faster, more cost-effective, and have facilitated the development of effective phage therapy (104). Similarly, PAS prediction programs could also be developed using bioinformatics. By utilizing accumulated big data, as in this study, deep learning could be applied to create predictive models for PAS, potentially leading to more efficient therapeutic strategies.
FUTURE RESEARCH DIRECTIONS
PAS has clear potential to become a revolutionary approach for treating antibiotic-resistant bacteria. However, research on this topic remains insufficient. Even with the data from the 83 studies analyzed in this review, the evidence is still too limited for the practical application of PAS as a therapeutic strategy. Therefore, further research on PAS is urgently needed.
Additionally, the same bacterial strains showed varying results with different antibiotics, and some antibiotics even exhibited antagonistic effects. This highlights the need for further investigation into these phenomena. It is also essential to conduct fundamental molecular biology studies to understand the mechanisms through which PAS operates with combinations of phages and antibiotics with diverse modes of action.
Given the vast number of phage types, consolidating all PAS-related data is a significant challenge. Moving forward, a more systematic organization of PAS studies will be necessary. Establishing systems such as a phage bank could facilitate research and clinical applications of PAS, making it a more accessible and reliable therapeutic option.




