INTRODUCTION
Norovirus is a highly contagious pathogen that causes acute gastroenteritis across all age groups, from infants and young children to adults (1). In the United States, approximately 2,500 foodborne norovirus cases are reported annually, while globally, norovirus is estimated to cause about 200,000 deaths and impose an economic burden of nearly USD 60 billion each year (2). The extensive genetic and antigenic diversity of noroviruses has posed a major obstacle to vaccine development. Based on sequence variation in the major capsid protein (VP1), noroviruses are classified into ten genogroups (GI–GX), which are further divided into multiple genotypes (2, 3). Of these, GI, GII, GIV, GVIII, and GIX are known to infect humans.
The viral genome comprises three open reading frames (ORFs). ORF1 encodes nonstructural proteins essential for replication, including RNA-dependent RNA polymerase (RdRp), helicase, VPg, and protease. ORF2 encodes the major capsid protein VP1, while ORF3 encodes the structural protein VP2, which functions as an auxiliary factor stabilizing the VP1 capsid (4). The current classification system is based on dual typing of the RdRp region of ORF1 and the VP1 gene, with 10 genogroups, 48 genotypes, and 60 P-types currently described (4, 5). Among these, GII.4 has historically been the predominant genotype responsible for global foodborne outbreaks. New GII.4 variants have emerged every 2–4 years, often linked to high regional or temporal prevalence. In particular, the GII.4 Sydney strain became globally dominant in 2012 (6, 7). However, in several Asian countries in 2014, GII.17 and GII.2 temporarily surpassed GII.4 as the predominant circulating strains (7, 8, 9, 10).
Due to its high transmissibility, broad age susceptibility, and propensity to cause large-scale outbreaks, norovirus places a substantial social and economic burden on public health systems and communities (11, 12, 13). Outbreaks are most commonly reported in daycare centers, kindergartens, schools, nursing homes, and hospitals, with transmission occurring through direct person-to-person contact, contaminated food, or waterborne exposure (14).
In this study, we investigated the epidemiological patterns of norovirus outbreaks associated with mass foodborne illness in food-service facilities in the Busan area. Our objective was to generate fundamental molecular epidemiological data to inform preventive measures against water- and foodborne infectious diseases and to support future vaccine development efforts.
MATERIALS AND METHODS
Sample Collection
This study investigated 61 outbreaks of foodborne illness that occurred in food-service facilities in the Busan area between January 2021 and November 2024. A total of 1,546 stool specimens were collected from symptomatic individuals and other persons epidemiologically linked to these outbreaks by local public health centers (Table 1). For the purposes of this study, food- service facilities were defined broadly to include institutions providing meals such as daycare centers, kindergartens, elementary, middle, and high schools, universities, and workplace cafeterias.
Table 1.
Number of outbreak and specimens collected from food-catering facilities in Busan, 2021–2024
Sample Pretreatment and Viral RNA Extraction
Approximately 1 g of each stool specimen was diluted in 9 mL of 0.1 M phosphate-buffered saline (PBS; Sigma-Aldrich, USA). Rectal swab specimens were suspended in 1.5 mL of sterile 0.1 M PBS. All samples were homogenized for 3 min using a vortex mixer and centrifuged at 6,000 rpm for 10 min at 4°C (1248R, Labogene, Korea). The resulting supernatants were used for nucleic acid extraction. Viral RNA was extracted using an automated nucleic acid extractor (Genolution, Korea). From each pretreated sample, 200 μL was processed with the NX-48 Viral NA Kit (Genolution, Korea), and the isolated RNA was subsequently used as a template for downstream molecular analyses.
Detection of Norovirus Genes
Norovirus detection was performed using real-time reverse transcription polymerase chain reaction (Real-time RT-PCR). The PowerChek™ Norovirus GI/GII Multiplex Real-time PCR Kit (Kogene Biotech, Korea) was used according to the manufacturer’s protocol and the guidelines of the Korea Disease Control and Prevention Agency (KDCA). Amplification was carried out on a thermal cycler (7500 Fast Real-Time PCR System, Applied Biosystems, USA), and samples were considered positive when norovirus RNA was detected by Real-time RT-PCR.
Genotyping and Phylogenetic Analysis
For genotyping, the RNA-dependent RNA polymerase (RdRp)–capsid junction region was amplified by conventional RT-PCR using a commercial one-step premix kit (iMOD-001TD, SnC, Seoul, Korea) with the primers listed in Table 2. The oligo nucleotide sequence was performed using forward primer to amplify the RdRp-ORF1 capsid region (15).
Table 2.
Primers used for RT-PCR of the Norovirus GI & GII detection
Amplification was performed on a thermal cycler (ProFlex PCR System, Applied Biosystems, USA) under the following conditions: reverse transcription at 42°C for 30 min; initial denaturation at 95°C for 15 min; 45 cycles of denaturation at 95°C for 1 min, annealing at 58°C for 1 min, and extension at 72°C for 1 min; followed by a final extension at 72°C for 10 min. Amplicons (570–579 bp) were verified using an automated electrophoresis system (QIAxcel Advanced System, QIAGEN, Germany) and subsequently sequenced by a commercial service provider (Macrogen, Korea). The obtained sequences were aligned with reference norovirus sequences deposited in GenBank, and genotypes were assigned based on the highest nucleotide sequence homology with Busan isolates.
Norovirus Genotype Identification and Homology Analysis
Sequences confirmed as norovirus were analyzed for genetic relatedness using MEGA version 12 (Molecular Evolutionary Genetics Analysis) (16). Multiple sequence alignments with previously reported strains were performed using Clustal W, and phylogenetic relationships were inferred by the Maximum Likelihood method with 1,000 bootstrap replicates. Genotypes of GI and GII strains were further identified using the NCBI BLAST search tool (http://blast.ncbi.nlm.nih.gov/Blast.cgi) and the Human Calicivirus Typing Tool (https://norovirus.ng.philab.cdc.gov/bctyping.html). Final classification was determined based on sequence homology with reference norovirus isolates deposited in GenBank (Table 3).
RESULTS
Incidence of Norovirus-Associated Foodborne Outbreaks
From January 2021 to November 2024, a total of 61 foodborne illness outbreaks were reported in food-service facilities in the Busan area, of which 41 (67.2%) were attributed to norovirus infection. The detailed incidence of norovirus-associated outbreaks is summarized in Table 3. When analyzed by facility type, daycare centers accounted for the highest number of outbreaks, followed by kindergartens and elementary schools, based on both outbreak frequency and the number of norovirus-positive samples.
Table 3.
Number of norovirus positive from food-catering facilities outbreaks, 2021-2024
| Daycare center | Kindergarten | Elementary school | Middle school | Military camp | Youth facility | Total | |
|---|---|---|---|---|---|---|---|
| 2021 | 71 | 38 | 4 | 9 | 39 | -* | 162 |
| 2022 | 59 | 48 | 13 | 15 | - | - | 135 |
| 2023 | 21 | - | 17 | 0 | - | 2 | 40 |
| 2024 | 4 | 6 | 34) | 13 | - | - | 57 |
| Total | 155 | 92 | 68 | 37 | 39 | 2 | 394 |
Norovirus Infection Prevalence by Outbreak Environment
The distribution of norovirus infections in food-catering facilities from 2021 to 2024 is presented in Fig. 1. Daycare centers were the most frequent sites of norovirus-associated foodborne illness, accounting for the highest number of outbreak cases as well as the largest proportion of genotyped isolates. Kindergartens and elementary schools followed in frequency.
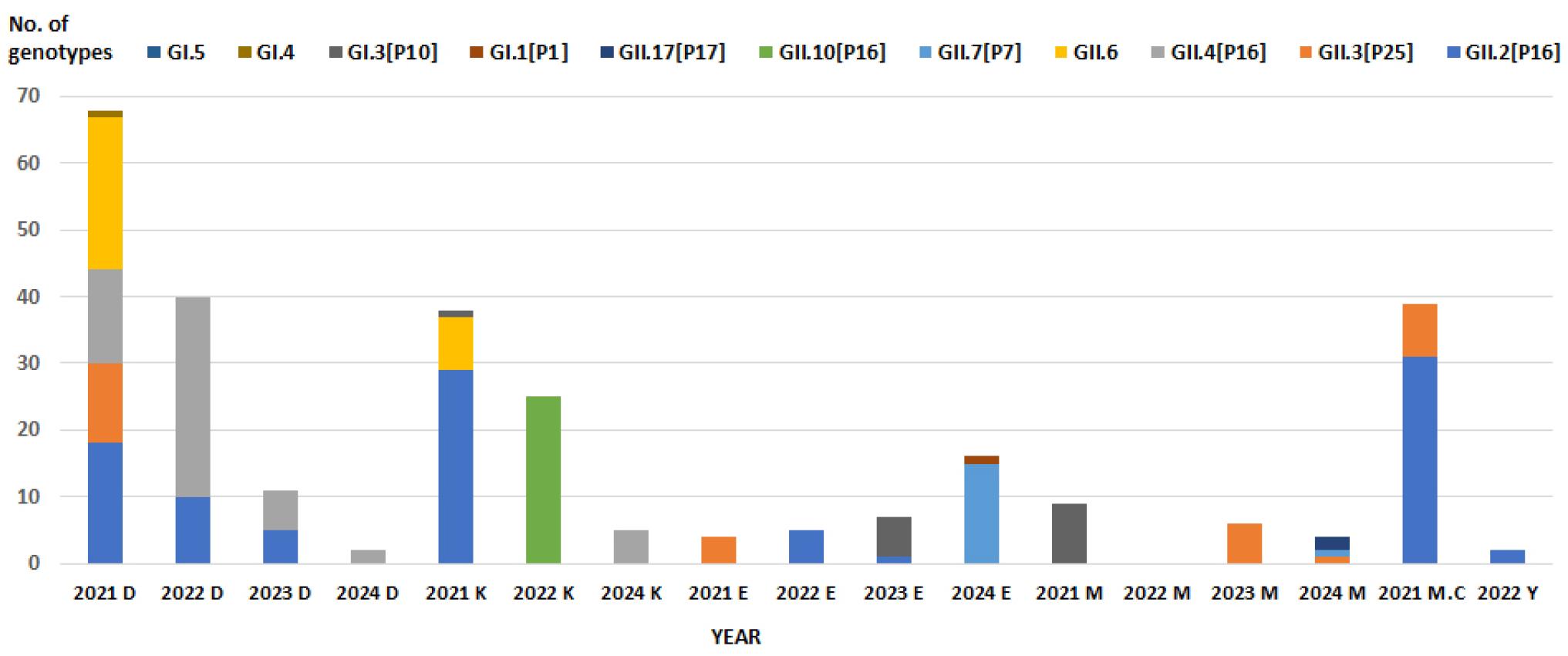
Fig. 1
Distribution of norovirus genotypes identified in food-catering facility outbreaks, Busan, 2021–2024.
Note that D, K, E, M, MC and Y denote daycare. kindergarten, elementary school, middle school, military camp and youth facility, respectively.
In daycare centers in Busan, the predominant norovirus genotypes varied annually; however, GII.2[P16], GII.3[P25], and GII.4[P16] were most frequently detected. Notably, analysis of outbreak patterns in daycare centers—the facilities with the highest number of foodborne illness cases—showed that the GII.4[P16] genotype circulated consistently each year during the study period (Table 4).
Table 4.
Norovirus-associated foodborne outbreaks in daycare centers, Busan, 2021–2024
In kindergartens, GII.2[P16] was the most frequently detected norovirus genotype, followed by GII.10[P16]. The predominant genotype varied annually, with GII.2[P16] in 2021, GII.10[P16] in 2022, and GII.4[P16] in 2024. No outbreaks were reported in 2023, limiting the ability to establish a consistent genotype pattern for kindergarten-associated outbreaks. Notably, GII.10[P16], which predominated in 2022, was not detected in 2023 or 2024, suggesting that its circulation represented a transient outbreak (Table 5).
Table 5.
Norovirus-associated foodborne outbreaks in kindergartens, Busan, 2021–2024
In elementary schools, norovirus-associated foodborne outbreaks occurred every year, with the highest number of cases reported in 2024. The predominant genotype shifted annually, being GII.3[P25] in 2021, GII.2[P16] in 2022, GI.3[P13] in 2023, and GII.7[P7] in 2024. Notably, GII.7[P7] was identified for the first time in 2024, marking its emergence during the study period (Table 6).
Table 6.
Norovirus-associated foodborne outbreaks in elementary schools, Busan, 2021–2024
In middle schools, foodborne outbreaks associated with meal provision were relatively infrequent compared with other food-service facilities. The predominant genotypes were GI.3[P13] in 2021, GII.3[P25] in 2022, and GII.17[P17] in 2024. Notably, GII.17[P17], which had first appeared in Korea about a decade ago (10) and was associated with multiple outbreaks at that time, re-emerged in 2024. This observation underscores the importance of sustained genotype-based surveillance (Table 7).
Table 7.
Norovirus-associated foodborne outbreaks in middle schools, Busan, 2021–2024
In other food-catering facilities, including high schools, youth centers, military camps, and hospital cafeterias, norovirus-associated foodborne outbreaks occurred only once or twice annually, making them less frequent than those in daycare centers and elementary schools. In a military base outbreak in 2021, both GII.2[P16] and GII.3[P25] were detected, indicating a mixed infection. Similarly, an outbreak at a youth center in 2023 was attributed to GII.2[P16].
Distribution of Norovirus Genotypes
Between 2021 and 2024, analysis of norovirus genotypes associated with foodborne outbreaks in food-service facilities revealed that, within the GI genogroup, more than half of the cases were attributable to GI.3[P10]. Within the GII genogroup, seven distinct genotypes were identified, among which GII.2[P16] and GII.4[P16] were the most prevalent (Fig. 2). These findings are consistent with previous reports: Hur et al. (2023) identified GI.3 as the predominant GI genotype and GII.4 as the predominant GII genotype in childcare centers in Gyeonggi Province, while Canon et al. (2021) reported that among children under five years of age, approximately 50% of norovirus infections were caused by GII.4, followed by GII.2, GII.3, and GII.6. The genotype distribution patterns observed in Busan thus parallel those reported in other regions and age groups (17, 18).
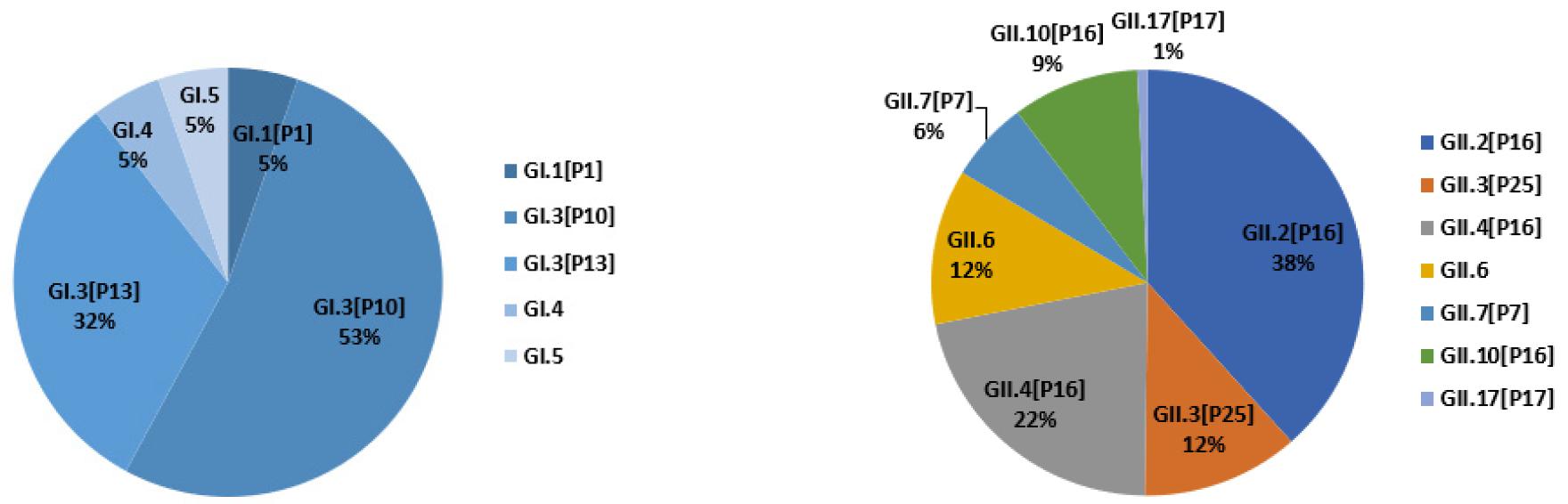
Fig. 2
Distribution of norovirus GI and GII genotypes identified in food-catering facility outbreaks in Busan, 2021–2024.
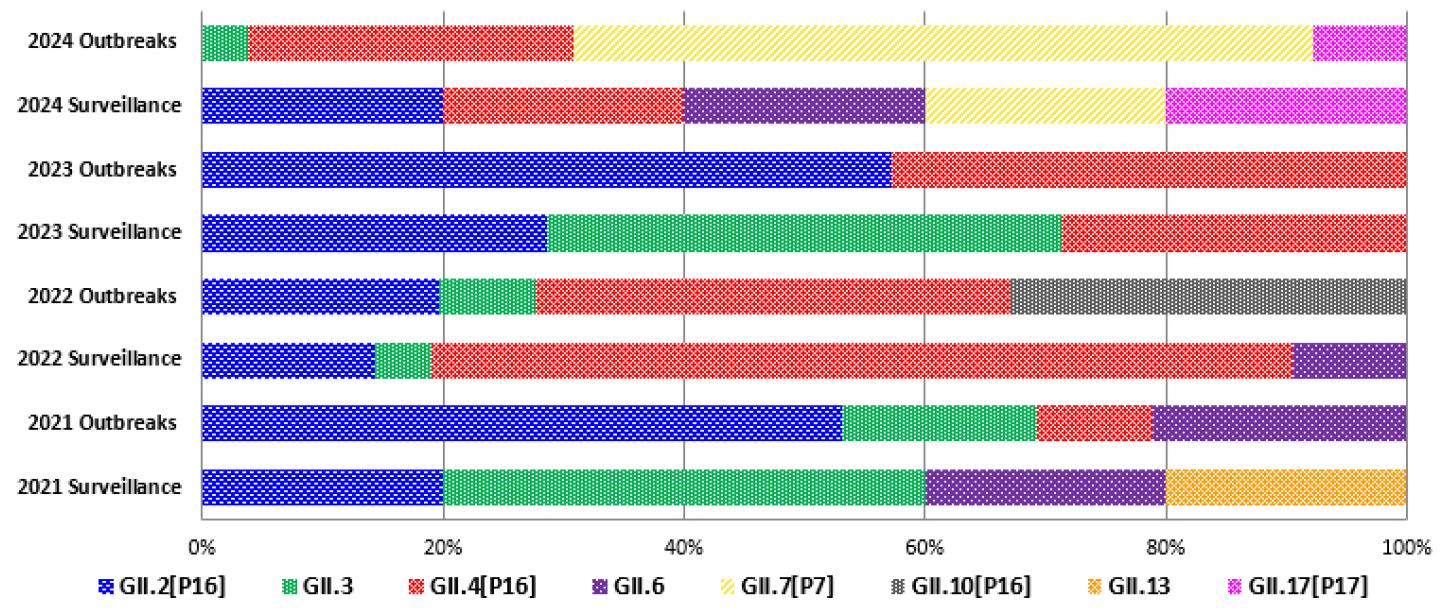
A comparison of norovirus genotypes detected in foodborne outbreaks at food-service facilities (2021–2024) and those identified in gastroenteritis patients visiting medical institutions is shown in Fig. 3. The annual distribution patterns were generally consistent between community outbreaks and hospital cases. Notably, the GII.7[P7] genotype, which emerged as the predominant strain in foodborne outbreaks in 2024, was also detected for the first time in gastroenteritis patients in Busan during the same year. In addition, the re-emergence of GII.17[P17]—previously associated with widespread outbreaks in Korea and Japan more than a decade ago—was confirmed (10, 19). These findings parallel U.S. CDC surveillance data, which identified GII.17[P17] as the most prevalent cause of norovirus outbreaks between September 2023 and October 2024 (20). Given the marked temporal fluctuations in norovirus epidemiology, sustained and systematic genotype surveillance will be critical for monitoring the emergence and spread of GII.7 and GII.17 variants.
Phylogenetic Analysis of Newly Emerging GII.7 and Re-emerging GII.17 Strains
A comparison of norovirus genotypes detected in foodborne outbreaks (2021–2024) and those identified in gastroenteritis patients at medical institutions is shown in Fig. 3. The annual distribution patterns were broadly consistent between community outbreaks and hospital cases. Notably, the GII.7[P7] genotype, which emerged for the first time in food-catering facility outbreaks in 2024 and became the predominant strain, was also identified for the first time in gastroenteritis patients in Busan during the same year. In addition, the GII.17[P17] genotype, which had caused widespread outbreaks in Korea and Japan more than a decade earlier, was found to have re-emerged. These results are consistent with U.S. CDC surveillance reports indicating that GII.17[P17] was the most prevalent cause of norovirus outbreaks between September 2023 and October 2024 (20). Given the substantial temporal variability of norovirus epidemics, continued genotype-based surveillance of GII.7 and GII.17 is critical for timely detection and response (Fig. 4).
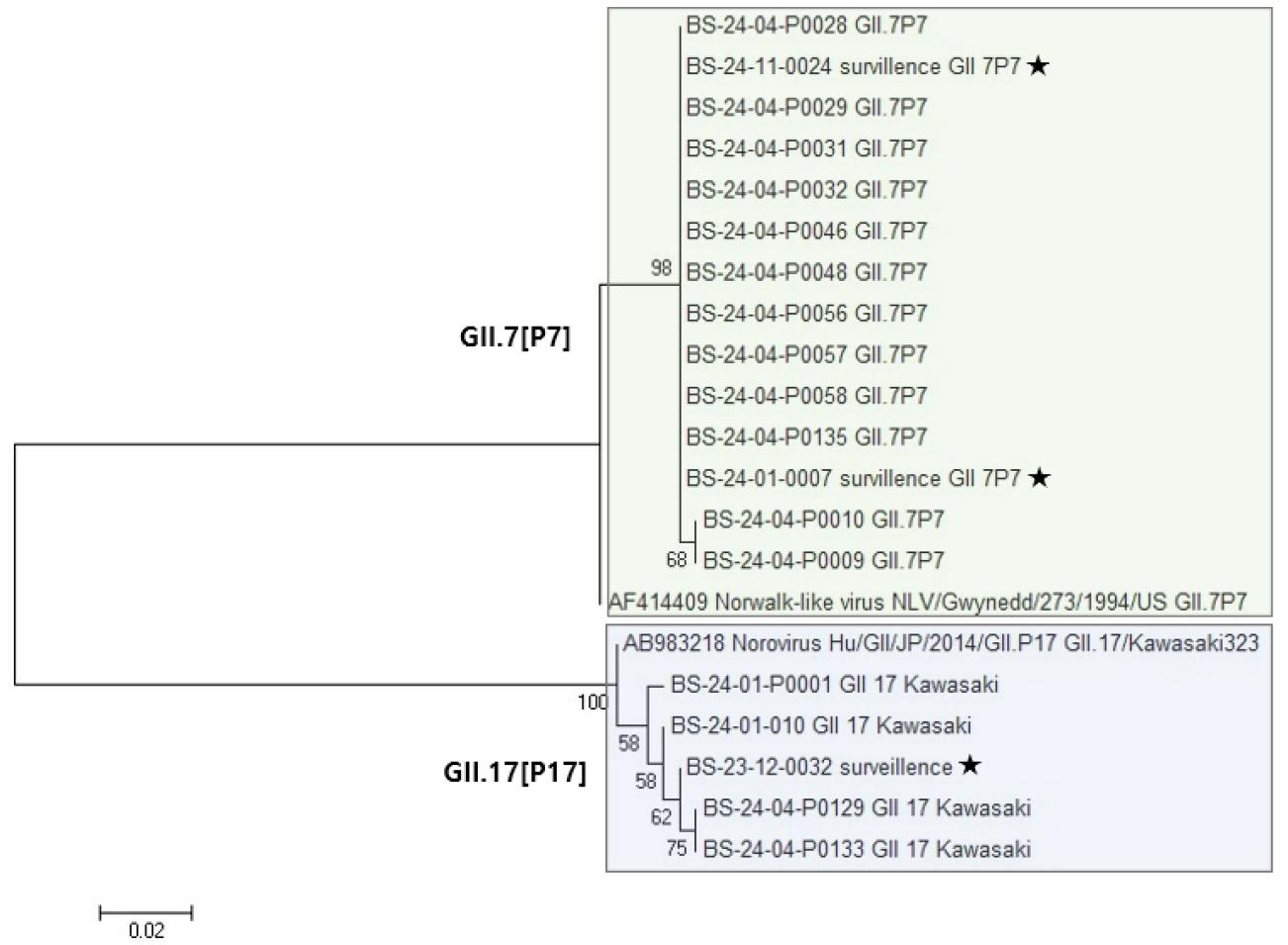
Fig. 4
Phylogenetic analysis of newly emerging norovirus GII.7 and re- emerging GII.17 strains based on the RdRp-VP1 region. The tree was constructed using Neighbor-joining method with 1,000 bootstrap replicates by MEGA 12. Square box indicates outbreak-associated strains detected over the four years and ★ symbols represent strains identified through laboratory surveillance of gastroenteritis patients.
DISCUSSION
Between January 2021 and November 2024, 61 foodborne-illness investigations were conducted in Busan. Norovirus was identified in roughly two-thirds of the outbreaks, yielding 394 laboratory-confirmed infections among 1,546 tested specimens. Norovirus testing was also performed on preserved food samples and on environmental swabs, including cooking utensils and door handles at food-catering facilities. None of these samples contained detectable viral RNA, illustrating a common limitation of epidemiologic fieldwork in tracing direct sources of norovirus infection. Although identifying the direct source proved difficult, the study systematically characterized norovirus outbreaks across a wide range of food-service facilities in Busan—including daycare centers, kindergartens, primary and secondary schools, youth training facilities, hospital cafeterias, and military bases over the four-year period. Annual surveillance revealed the circulation of multiple genotypes, and homology analysis with reference strains enabled the identification of facility-specific genotype distributions.
No distinct outbreak patterns emerged to distinguish among different types of food-service facilities. However, daycare centers consistently exhibited the GII.4[P16] genotype throughout the study period. Because daycare facilities serve preschool children—a group with relatively immature immunity—this finding underscores the critical importance of targeted prevention strategies in such settings. In addition, daycare environments involve intense person-to-person contact and the sharing of toys, conditions that are highly conducive to persistent transmission of norovirus. From a public health perspective, the persistent circulation of GII.4 in young children highlights the urgent need for vaccine development against this genotype, which may serve as a cornerstone for reducing transmission in vulnerable populations. Consistent with our findings, Hur et al. (2023) reported that GII.4 was also dominant in childcare centers in Gyeonggi Province (17).
In contrast, foodborne outbreaks associated with elementary and middle schools exhibited greater genotype variability, with no detection of GII.4 during the past four years. Instead, different genotypes predominated each year, and in 2024, the newly emerging GII.7 strain became dominant. Because school-aged populations engage in broader social interactions through mixed classes and extracurricular activities, opportunities for the introduction and spread of novel or reemerging genotypes are increased. This genotype was also detected through the national water- and foodborne pathogen surveillance program targeting acute diarrhea patients, suggesting itsgenog broader circulation beyond school-associated outbreaks (21, 22).
There is a genotype shift of norovirus outbreaks in the United States Between 2022 and 2025 with emerging GII.17 with a substantial decrease in GII.4 which was the predominant causes of US norovirus outbreaks (23). Novazzi et al (2025) described a large GII.17[P17] outbreak in a school complex in northern Italy (24).
The dynamic genotype turnover suggests that contact environments and host immunity are major determinants of norovirus transmission. Immunity to norovirus is known to be incomplete and of limited duration. Children attending daycare centers experience limited exposure to various norovirus genotypes. This immunologic immaturity allows repeated infections with norovirus genotypes within short intervals because of limited immunity among genotype (25, 26).
Collectively, these findings illustrate the complex and evolving nature of norovirus epidemiology in community-based food-service facilities. Continuous genotype-based monitoring is essential not only for early detection of newly emerging genotypes such as GII.7 but also for tracking the persistent presence of strains like GII.4 in susceptible populations. Strengthened genomic and environmental monitoring will be essential to detect, interpret, and mitigate the next shift in norovirus genotype dominance. Ultimately, such surveillance provides a foundation for more effective outbreak prevention, improved management of acute gastroenteritis, and the development of evidence-based vaccination strategies.